Watch Episode Here
Listen to Episode Here
Show Notes
Michael Levin joins the Stanford Chemical Engineering Colloquium to explain how non-neural bioelectricity lets cell networks make decisions about body shape and regeneration. He describes an “anatomical compiler” for telling cells what to build, shows real multi-headed and multi-limbed animals as proof of concept, and explores how electroceuticals could transform birth defect repair, limb regrowth, cancer control, and synthetic bioengineering.
CHAPTERS:
(00:00) Overview and main claims
(02:09) Anatomical compiler and specification
(06:29) Regeneration and inverse problems
(11:47) Robust morphogenesis and computation
(17:50) Non-neural bioelectricity fundamentals
(24:38) Rewriting pattern with bioelectricity
(32:05) Bioelectric memory and setpoints
(38:27) Bioelectric regeneration and translation
(42:56) Multi-scale modeling and simulation
(46:29) Bioelectric control of cancer
(49:40) Designing electroceutical interventions
(53:27) Summary, implications, and acknowledgments
PRODUCED BY:
SOCIAL LINKS:
Podcast Website: https://thoughtforms-life.aipodcast.ing
YouTube: https://www.youtube.com/channel/UC3pVafx6EZqXVI2V_Efu2uw
Apple Podcasts: https://podcasts.apple.com/us/podcast/thoughtforms-life/id1805908099
Spotify: https://open.spotify.com/show/7JCmtoeH53neYyZeOZ6ym5
Twitter: https://x.com/drmichaellevin
Blog: https://thoughtforms.life
The Levin Lab: https://drmichaellevin.org
Lecture Companion (PDF)
Download a formatted PDF that pairs each slide with the aligned spoken transcript from the lecture.
📄 Download Lecture Companion PDF
Transcript
This transcript is automatically generated; we strive for accuracy, but errors in wording or speaker identification may occur. Please verify key details when needed.
Slide 1/54 · 00m:00s

Thank you for the opportunity to speak to all of you and to share some work and get your feedback. If anyone would like more details or to get in touch with me, all the papers and everything else are at these websites.
There are four basic points that I would like to get across today. The first is that there are really some very significant knowledge gaps about the control of large-scale anatomical homeostasis. It encompasses our understanding of the relationship between the genome and the anatomy. I'll show you why things are quite puzzling there. I will claim that fundamental advances in new biomedicine are going to require not only understanding the molecular mechanisms for this process, but also the decision-making by cellular collectives that is sufficient for it to happen. That a key medium for computation in living tissue is non-neural bioelectricity. We've developed some techniques to manipulate this layer of physiological software, and I'll show you why I say that. Fundamentally, looking towards the future, I will argue that cracking this bioelectric code will enable a novel approach using electroceuticals for applications in birth defects, regenerative medicine, cancer, and synthetic bioengineering.
Slide 2/54 · 01m:28s
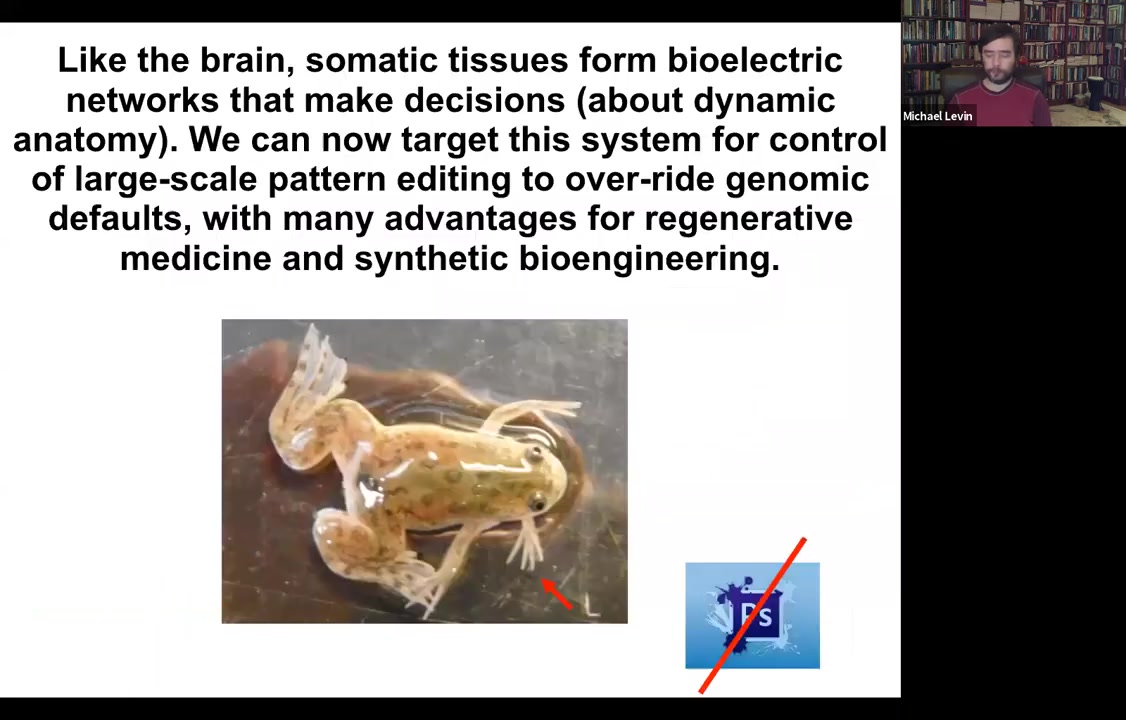
Boiling down the whole talk into two sentences, what I'm going to tell you is that your body tissues form electrical networks that make decisions. These are decisions about dynamic anatomy, and we now have the ability to target the system to control large-scale editing that can even override genomic default states with advantages and opportunities for regenerative medicine and synthetic bioengineering. I show you one of our five-legged frogs to point out that you're going to see weird creatures today. None of this is Photoshop. These are all actual living things that represent our attempt to test some of the models we have.
Let's look at these knowledge gaps.
Slide 3/54 · 02m:13s
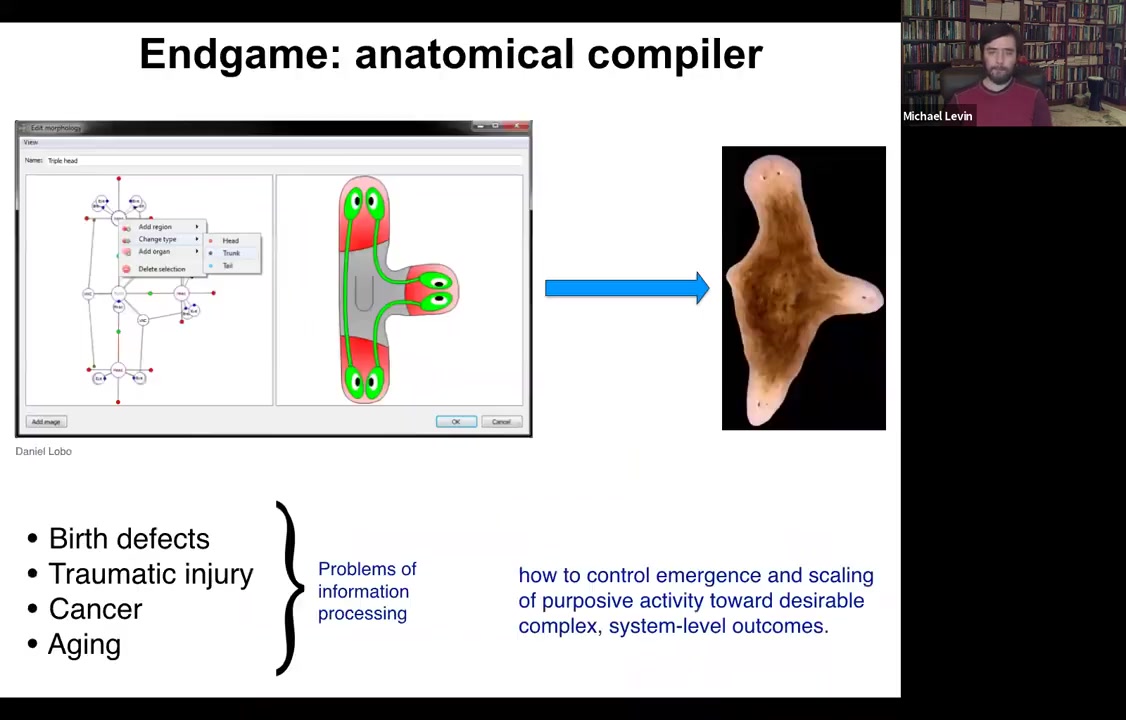
If we think about what is the end game of our field, where are we going? At what point can we all go home? What we would like to have is something that we call the anatomical compiler.
The deal is that you ought to be able to sit down and draw the animal or plant that you would like to have at the level of the anatomy. Not at the level of pathways, but at the level of final anatomy, the way that we do with machine parts and things like this. You would be able to draw this three-headed worm. If we knew what we were doing, we would have the capability of having the software, which would then compile this anatomical description down into a set of stimuli that would have to be given to cells that would cause them to produce whatever it was that you just drew.
Here's this three-headed flatworm. Now, the reason that this is fundamentally important is that almost all of the problems of biomedicine — except for infectious disease, pretty much everything else: birth defects, traumatic injury, cancer, aging — would be solved if we had the ability to tell cells what to build. We would need the knowledge to control what cellular collectives cooperate towards building.
All of these things are in some important sense problems of information processing. How is it that cells work together to build these things? We are very far from having anything like this, and only in a few very special cases do we know anything about how to make specific shapes come out. Let's think about why that is.
Slide 4/54 · 03m:52s
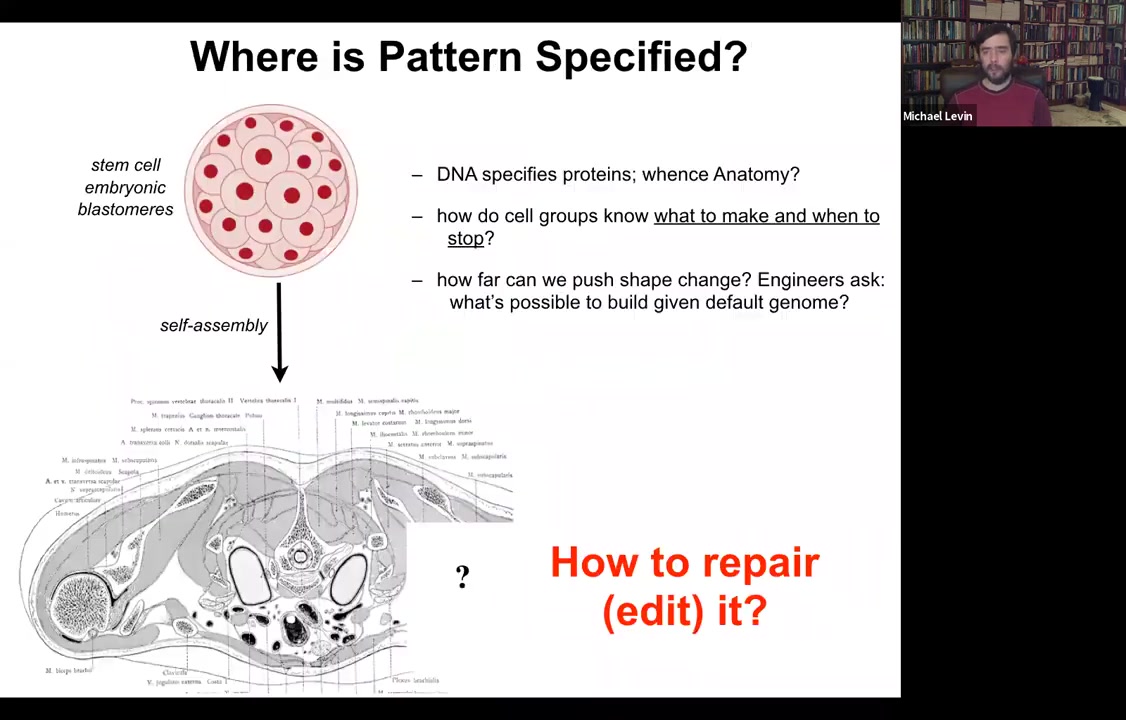
Where is anatomy specified?
We all start life roughly like this. This is a collection of embryonic blastomeres. Shortly thereafter, you get something like this. This is a cross-section through a human torso.
Look at this incredible invariant order.
All of these organs are the correct shape, size, position, orientation.
Slide 5/54 · 04m:16s
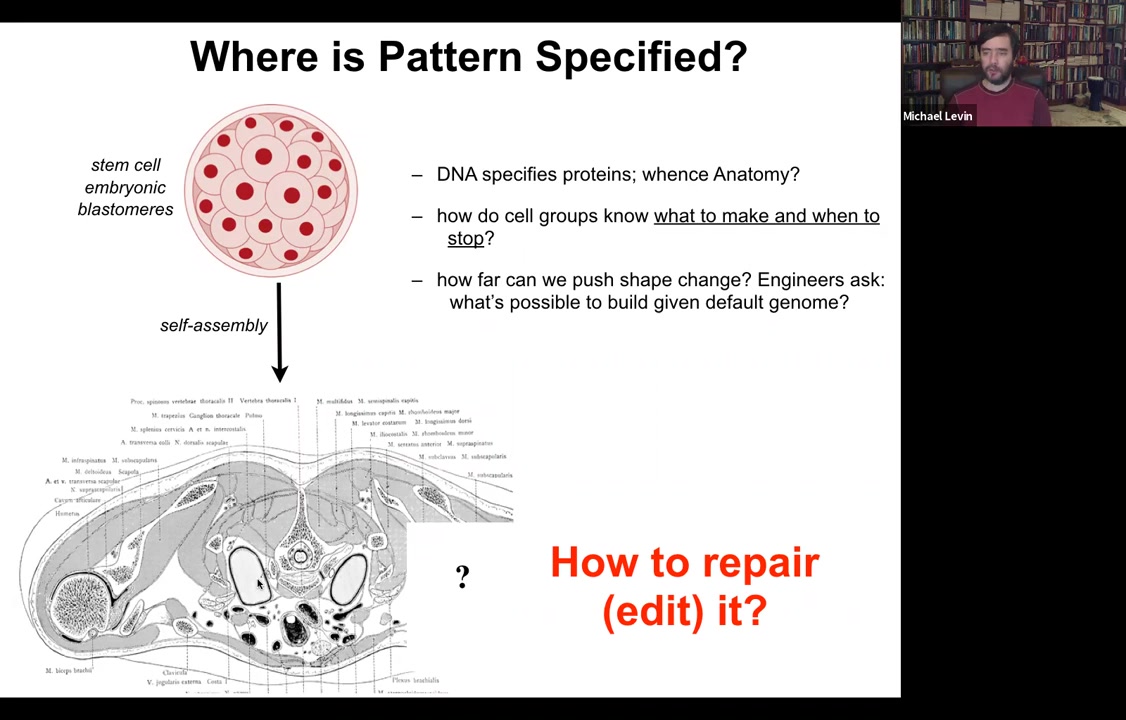
Everything is exactly the right stuff. It's absolutely a staggering amount of order. We would like to know, where is this information coming from? We're tempted to say DNA, but of course we know what genomes specify. The DNA specifies proteins. There's nothing directly in there that specifies any of this. We would like to know how cells use the genomically specified hardware that they have to know what to make and when to stop. In regenerative medicine, if a piece of this is missing, how do we get the cells to rebuild it?
As engineers, we ask what's actually possible to build given any particular default genome. The reason that in my group we frame this thing as a collective intelligence problem is that individual cells are extremely competent.
Slide 6/54 · 05m:02s
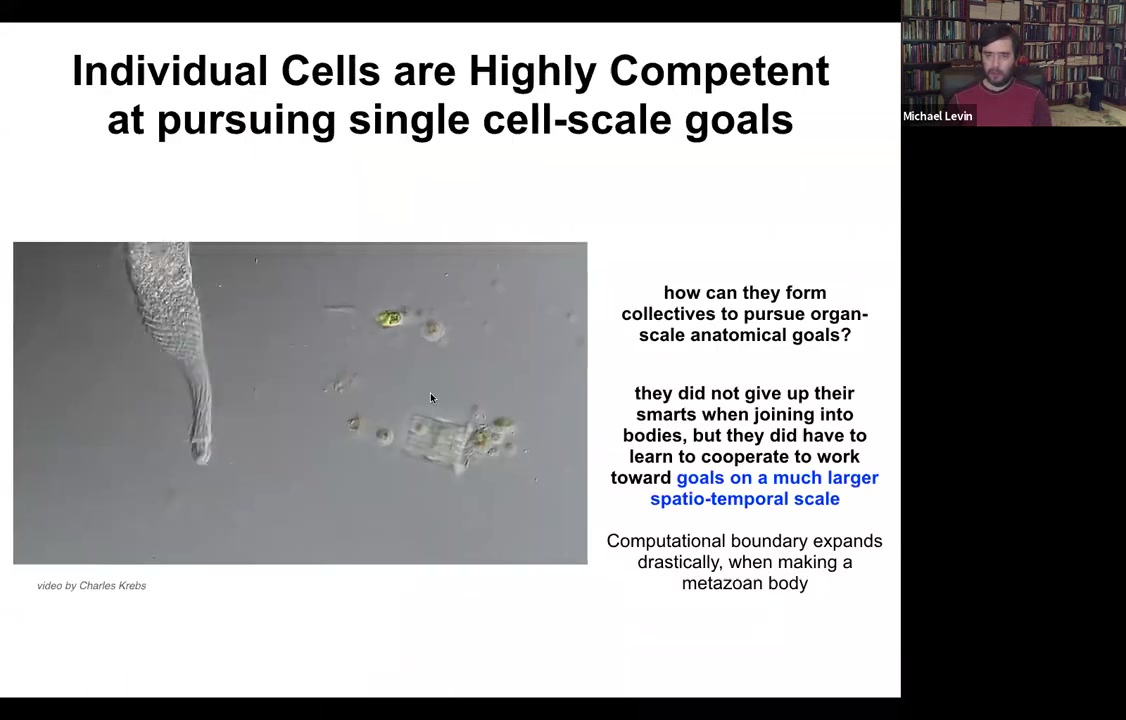
Here you see one cell. This is a single-celled organism. This is called Lacrymaria. There is no brain, there's no nervous system, there are no stem cells, there's no cell-to-cell communication, just one cell handling all of its local goals. It's handling its physiological goals, anatomical control, metabolic, behavioral, everything is handled by this one cell in real time. The amazing thing is that these cells, which are extremely competent in their local, very small environment, when they work together to make a metazoan body, they can work on much bigger goals.
Slide 7/54 · 05m:41s
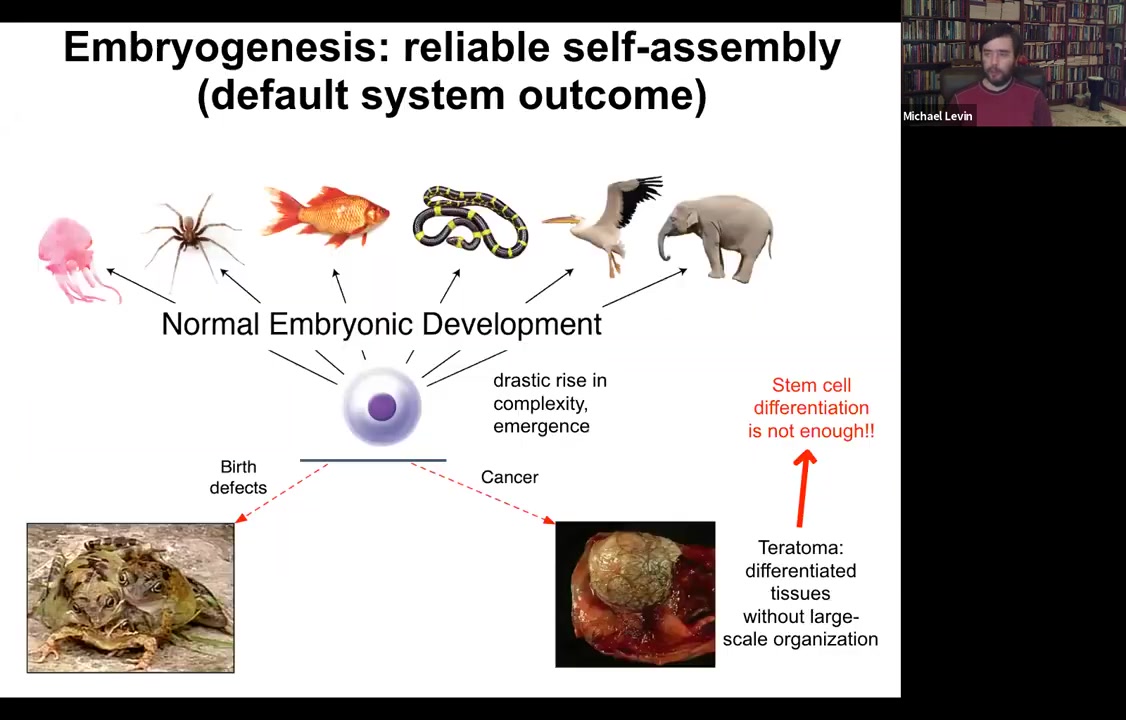
What you see is an inflation, not only of body size, but actually of their ability to pursue states in various kinds of problem spaces, including anatomical morphospace, physiological space.
A single cell can give rise to a collection of cells that self-assembles into some incredibly complex morphology.
We know that simply understanding stem cell biology is not going to be enough because here we have a teratoma and this thing might have hair and teeth and bone and muscle and skin. The work of the stem cells has proceeded fine. You have all your derivatives, you have your various tissues. What you don't have is this three-dimensional structure. We need to understand how this works.
Slide 8/54 · 06m:29s
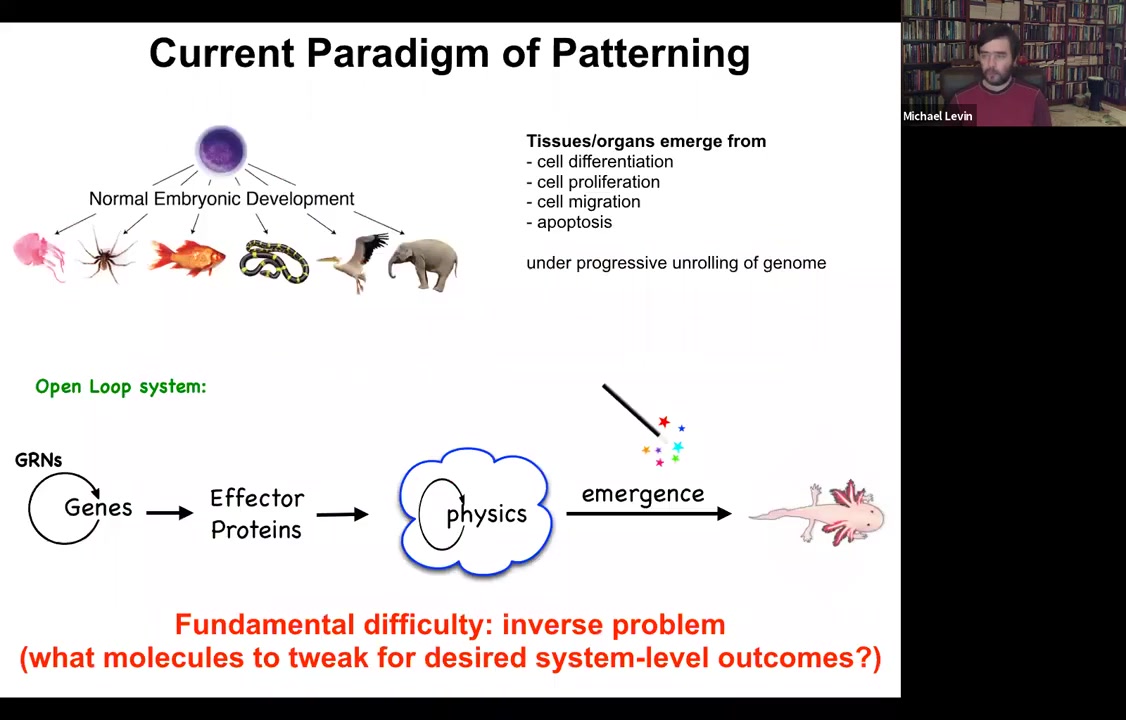
In standard developmental biology, the story that we're told is a kind of feed-forward open-loop system, which is very much based around emergence. The idea is that there are gene regulatory networks, so genes turn each other on and off. Some of these genes are effector genes; they code for effector proteins, so they are sticky or they diffuse or they exert force or something like this. Then there's this physical process where all of these things interact with each other. Then through things that are studied by the science of complexity, out come these amazingly complex results. Like the salamander here.
This story is certainly true in the sense that these things all happen. But it's incomplete, and it has a fundamental difficulty, which is this inverse problem. If we're committed to this feed-forward emergence story, then making changes here requires us to exert our interventions, for example, at the genetic level. That means that we have to try to invert this process of emergence, which is fundamentally insolvable. Most of these inverse problems are too difficult to solve. How do you know what genes to tweak to make desired changes at the three-dimensional anatomy level?
Slide 9/54 · 07m:50s
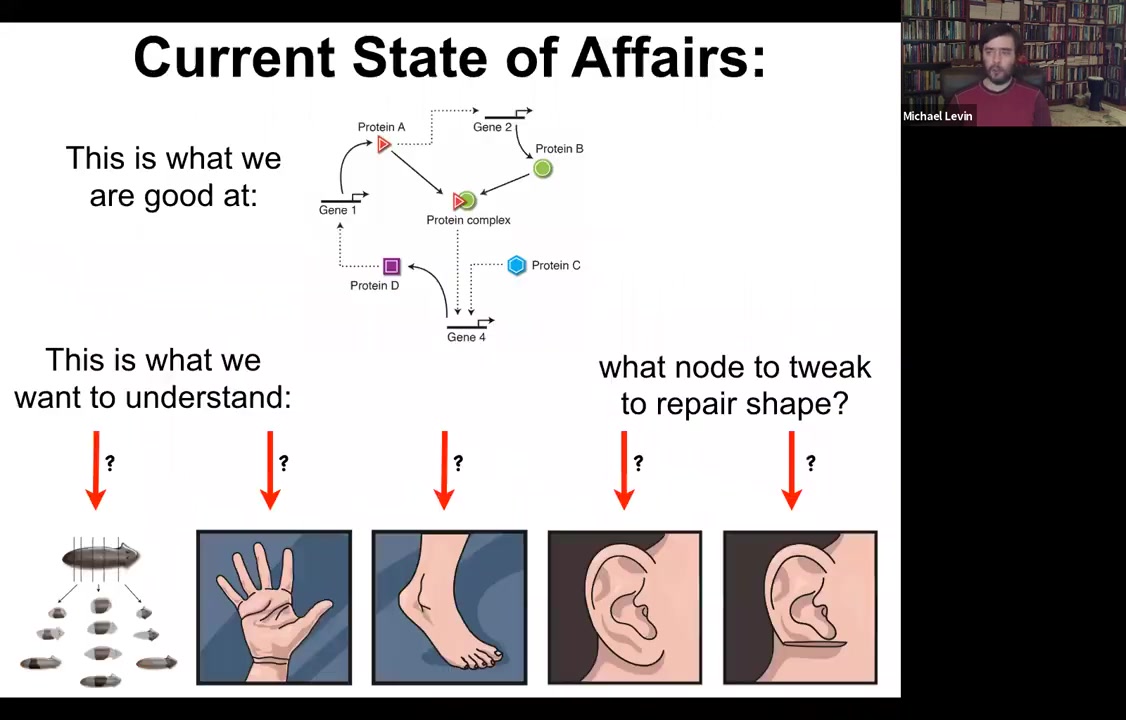
The current state of affairs is this. We're very good at figuring out the lowest hardware level of which genes and proteins interact with which other genes and proteins. But what we'd really like to understand are things like this: the ability of planaria to regenerate from pieces, the shape of the hand and the shape of the foot and why they're different, and what you would have to do to get these specific shapes back. It's not obvious what you would do at this level.
Slide 10/54 · 08m:18s
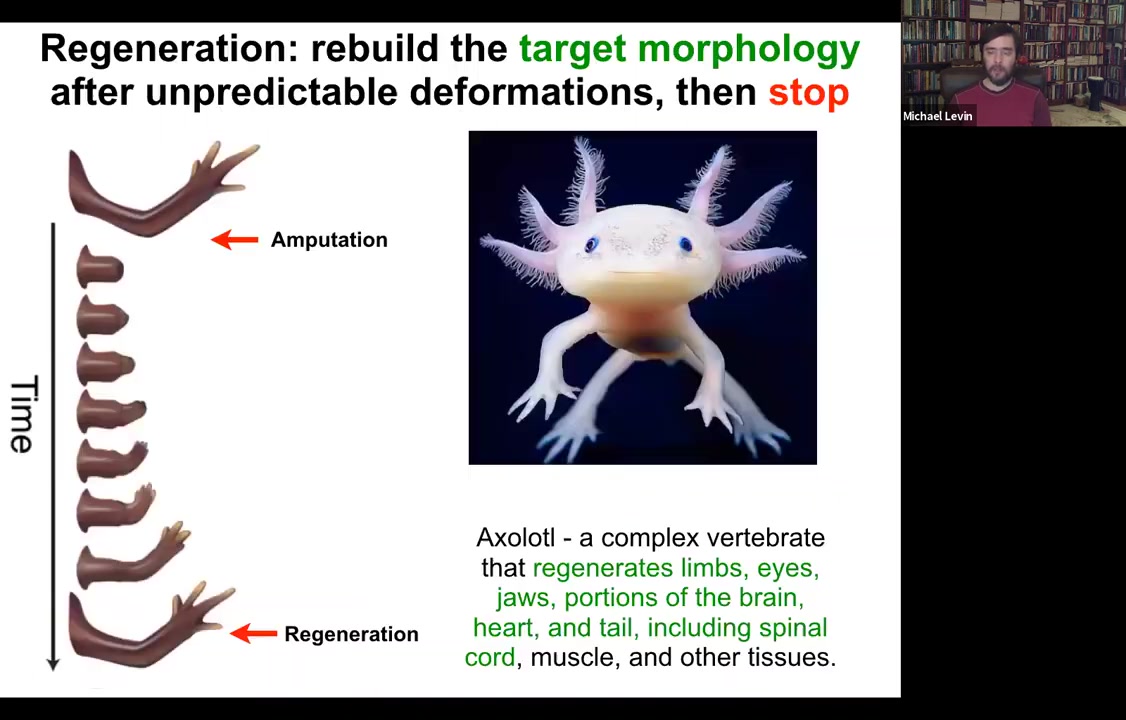
So the amazing thing is that... Some animals are very good at this, and their bodies show remarkable plasticity in this kind of collective activity of the cells. So for example, here, this animal is a salamander. It regenerates its limbs, its eyes, its jaws, its portions of the brain and the heart. So if they're amputated, they will grow back. And What's cool is that not only is this process incredibly flexible, meaning if you amputate at the shoulder, you grow the whole thing. If you amputate at the wrist, you start here and you just grow the parts you need, but then it stops. And that's the most remarkable part of all of this. And so lots of people work on trying to kickstart regeneration. But actually, how does it know when to stop? Because when it stops is when it has built a correct salamander arm. That's when it stops. How does the system know when it's built a correct salamander arm?
Slide 11/54 · 09m:10s
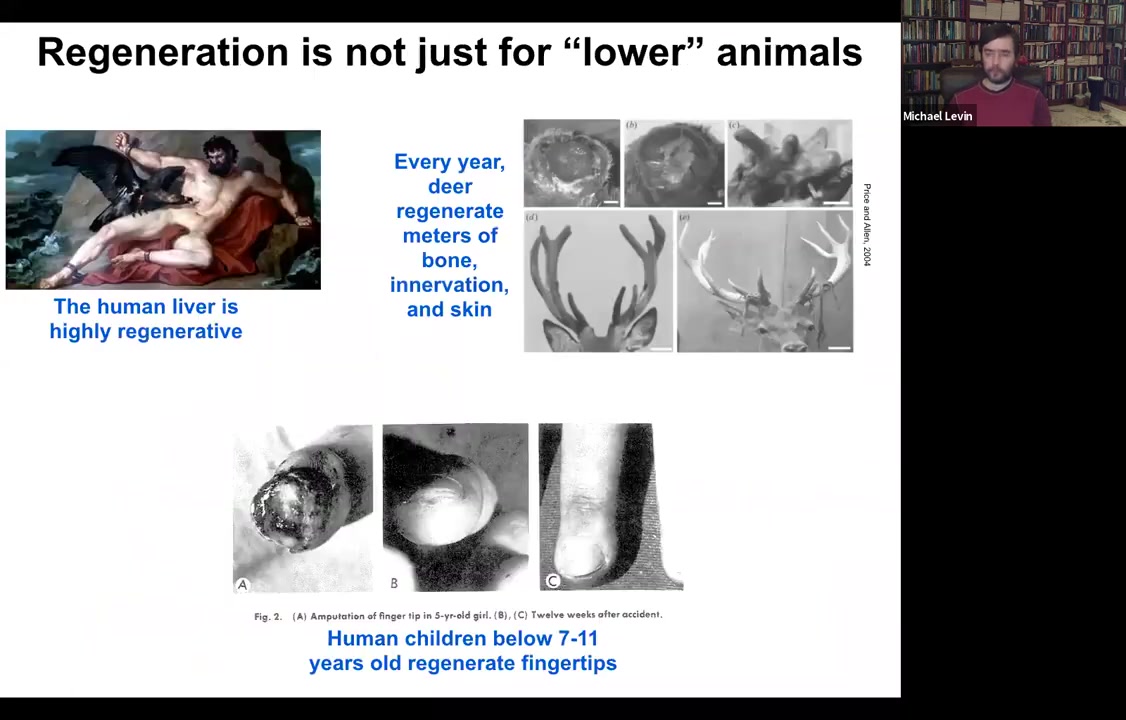
It's important to note that this is not just some weird quirk of salamanders. Humans and other mammals can do some of this. The human liver is highly regenerative. That's been known for a long time. Unclear to me how the ancient Greeks knew that, but they clearly did. Deer regenerate huge amounts of bone, up to a centimeter and a half of new bone per day. Bone, vasculature, innervation, and skin can regenerate. Even human children can regrow fingertips. It usually stops at a particular age, but if you keep it clean, the amputation will give rise to a cosmetically perfect finger.
Slide 12/54 · 09m:51s
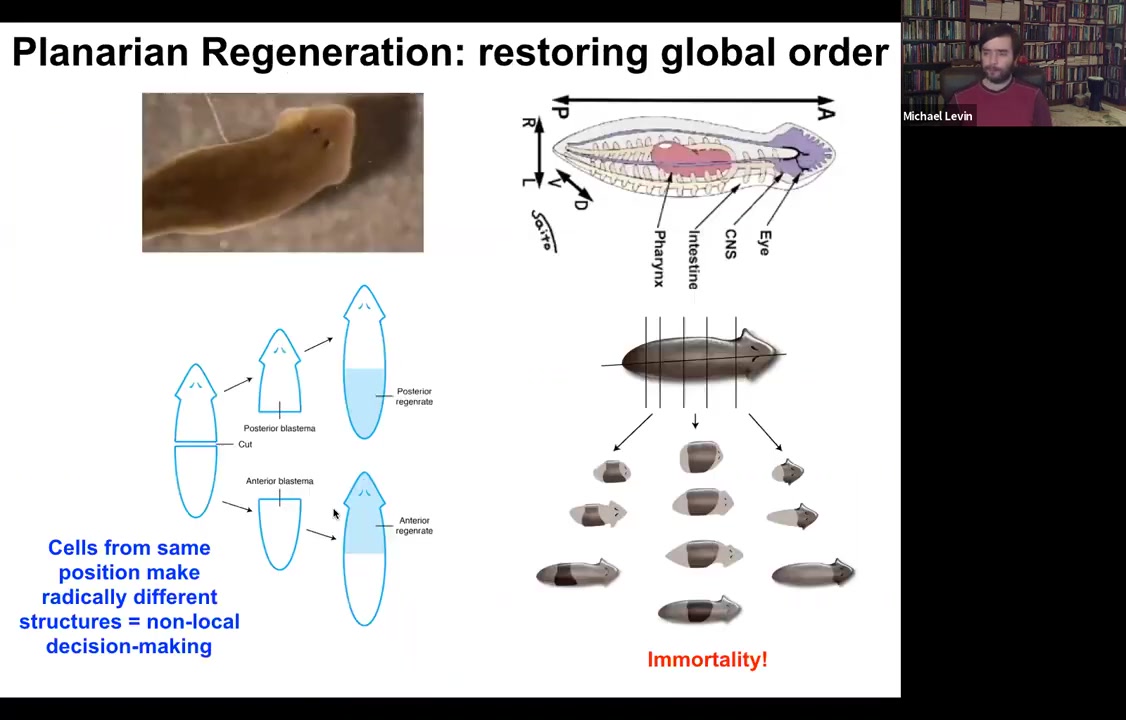
So the champions of this process are these guys. These are planaria. These are flatworms. They have a true brain, central nervous system, the same neurotransmitters that you and I have. And the amazing thing about them is that you can cut them into lots of pieces. The record is 275. And every piece will regrow exactly what's missing, no more, no less, to give you a perfect tiny little worm.
While the new stuff is growing, the remaining tissue is shrinking so that they will all, as quickly as possible, get to proportion, to correct proportionality. And the other thing about them is they're immortal. There's no such thing as an old planarian. So if you're interested in aging or these ideas that things inevitably wind down and accumulate errors and so on, planaria are telling us that is not absolutely required. Here's a life form that's basically immortal.
We're really not very far along in understanding how the cellular collectives make decisions.
Slide 13/54 · 10m:57s
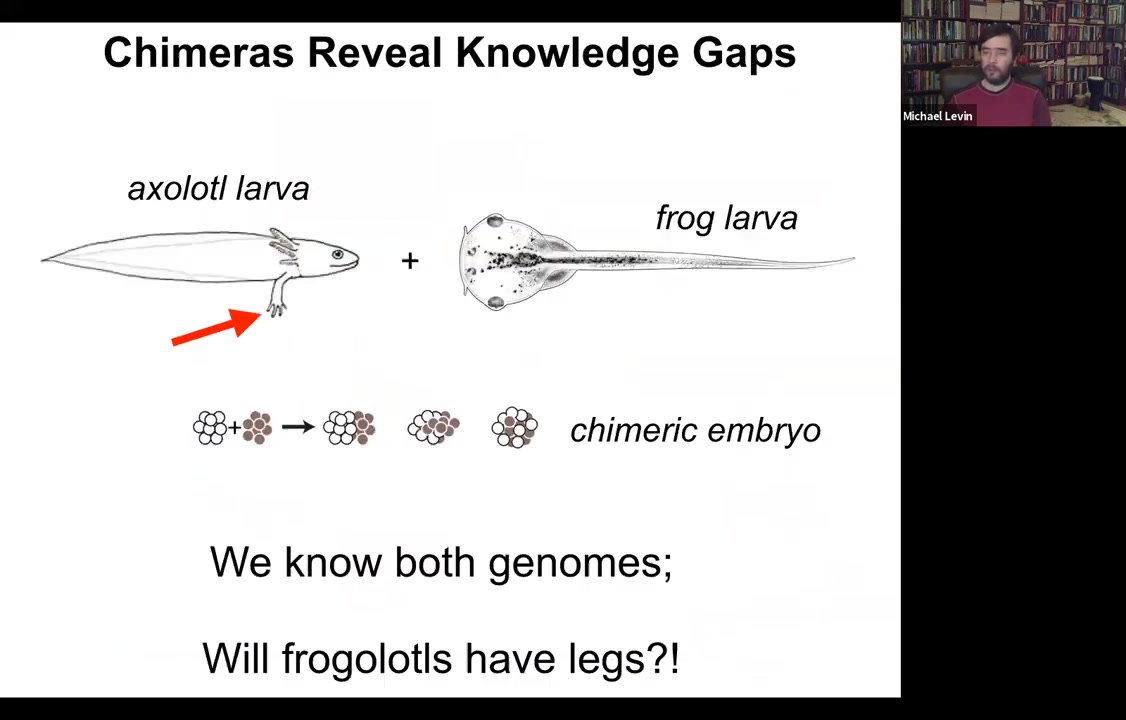
One of the easiest ways to see that is to think about chimeric organisms. Here's a simple example, which we're making in my lab right now. Here's an axolotl larva. Axolotl larvae have legs. Here's a frog larva, a tadpole. Tadpoles do not have legs.
One can ask a simple question. If I combine early axolotl tissue with early frog tissue in early embryogenesis, I make a chimeric embryo. They're perfectly healthy. We call them frogolotls. We have the genomes, we have the axolotl genome, we have the frog genome. Now we ask a simple question: do frogolotls have legs? Even though we have all this information, we have no idea how to predict in advance whether frogolotls will have legs. If so, will they be made entirely of axolotl cells or of both types of cells? These are the kinds of things that we would like to understand.
Slide 14/54 · 11m:47s
One thing that's very important as part of this process is specifically the algorithm that enables robustness and the handling of novelty. I'll show you a simple example of that. We discovered this a few years ago.
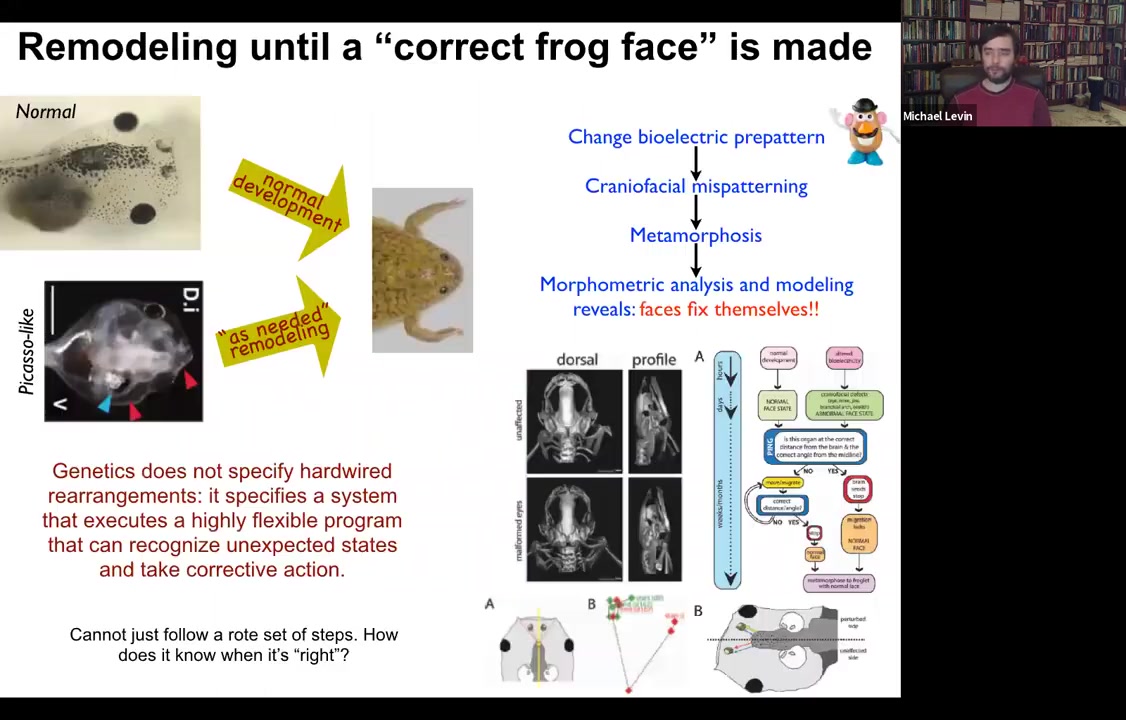
Here is a tadpole. It has eyes here, it has nostrils and a mouth down here. All of these things have to move around in order to get to a frog face. During metamorphosis, the face has to deform. The jaws have to come out, the eyes have to move forward, everything has to move. It was thought that somehow what the genome did was to give each piece of the face a particular direction and amount of movement. That way standard templates become standard frogs.
What we did was we created what we call Picasso tadpoles. Everything's in the wrong position. The eyes are on top of the head, the mouth is off to the side, everything is mixed up. I'll show you in a minute how we do it. But the amazing thing is that these animals become pretty much normal frogs because all of these different organs move around through unnatural paths and sometimes they go too far and actually have to back up.
Slide 15/54 · 12m:53s
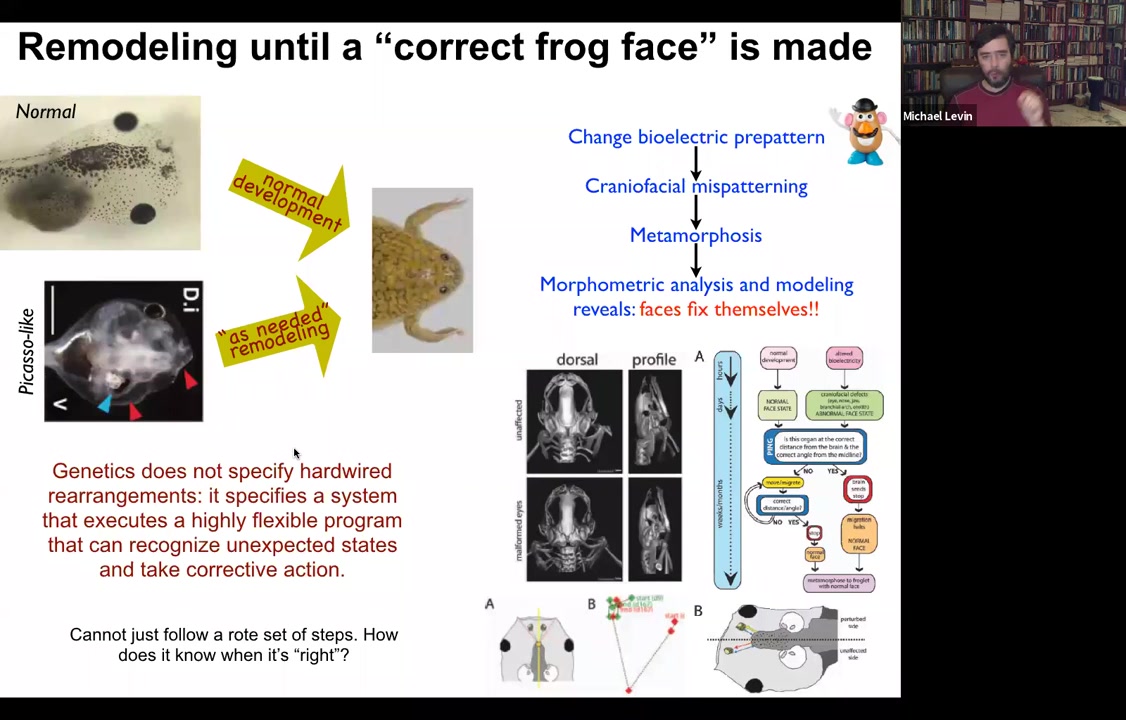
But everything moves around until it gets to a correct frog face and then the remodeling stops. So in fact, what the genetics specify is not a bunch of hardwired rearrangements, but a system that executes a really flexible error minimization scheme. It's able to start off at incorrect or abnormal positions and still get to where it needs to be. This parenthetically matches William James's definition of intelligence, which is the ability to reach the same outcome despite perturbations and starting from novel starting configurations. So how does this system know what a correct face is, and how do we get there?
Slide 16/54 · 13m:31s
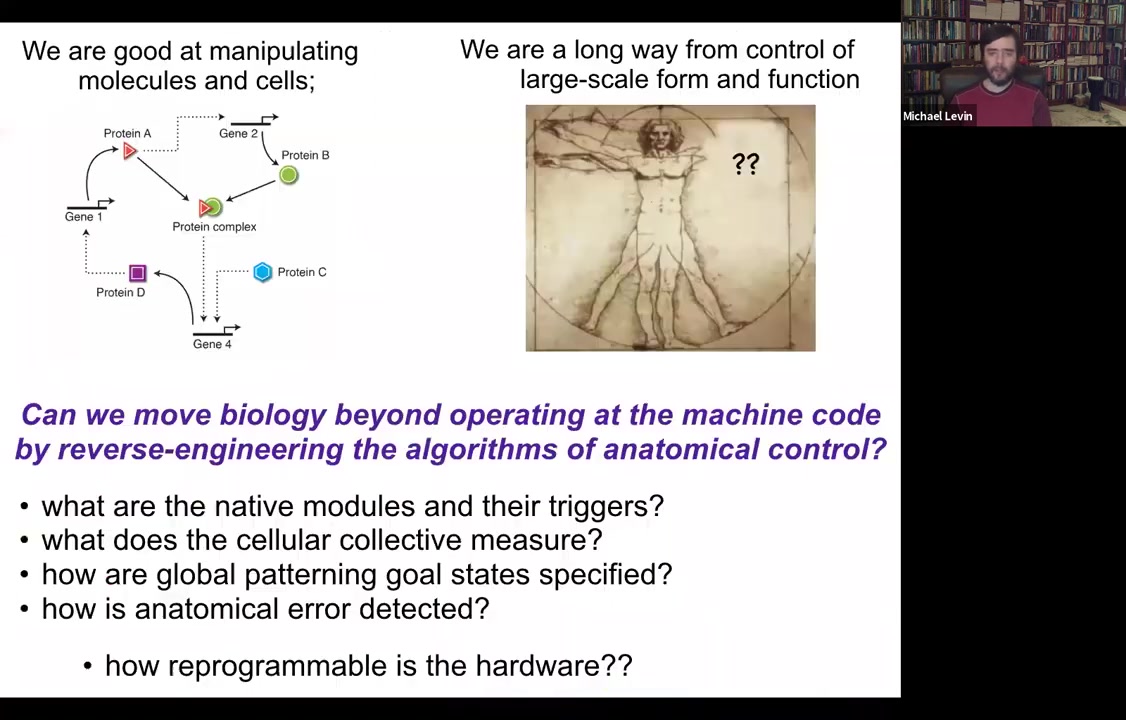
So we've been thinking about this problem. My background is computer science. And so to me, all of this looks like a problem of information processing at different levels. And what I would love to know is: could we go beyond the hardware and operating at this level and ask about the algorithms? How are these decisions being made? What do the cellular collectives measure? What are their modules or subroutines? How are these global patterning goals specified and stored? And in particular, how reprogrammable is any piece of biological hardware?
Slide 17/54 · 14m:07s
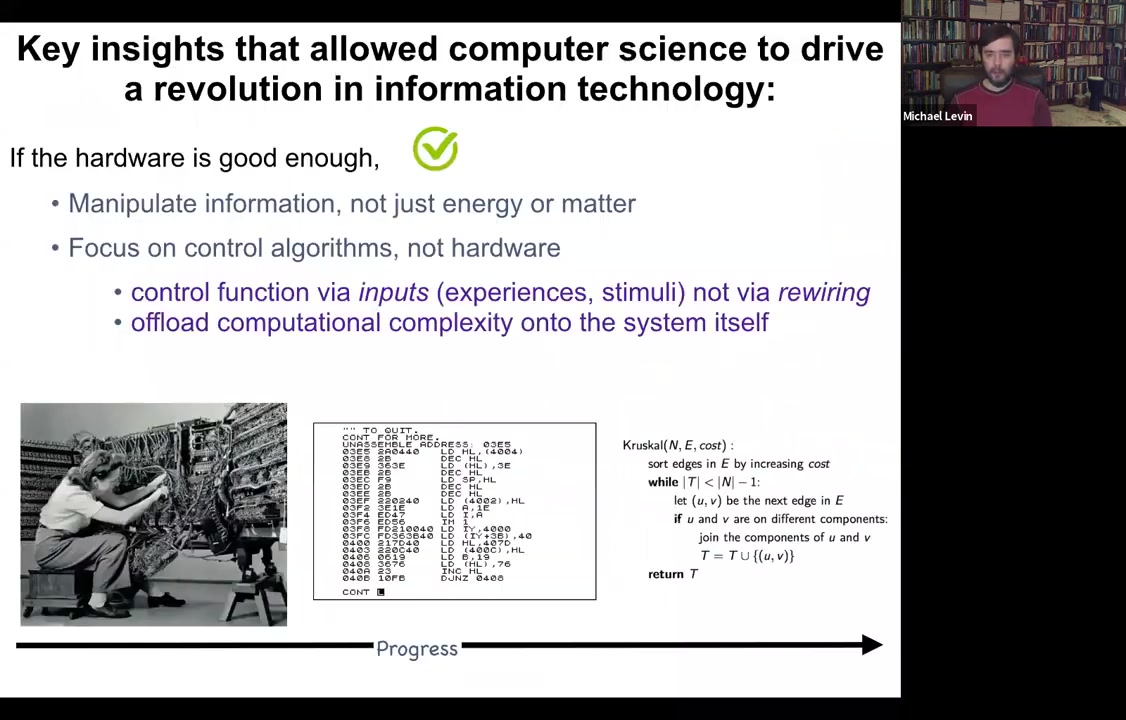

So in computer science, there was this really interesting journey that we took from, this is what programming looked like in the 1940s and 50s. In order to get the computer to do something different, you physically had to rewire it. You had to rearrange the hardware. You had to move the wires. But what computer scientists realized is that if your hardware is good enough, and I'm going to argue that biological hardware is definitely good enough, what you can do is something really interesting. You can reprogram the machine without touching the hardware, by giving it stimuli or experiences, inputs. And that will allow you to program in a much higher level language where you're not drowned in the details of the hardware or even the machine language, but you're actually able to take advantage of some really high level types of programming strategies to get very complex kinds of outcomes. And I'm certainly not arguing that living things are computers in the sense that they're not built to the same architectures that our current computers are. But this idea of reprogrammable hardware and of controlling a system by inputs and stimuli as opposed to physical rewiring, it's very powerful.
Slide 18/54 · 15m:15s
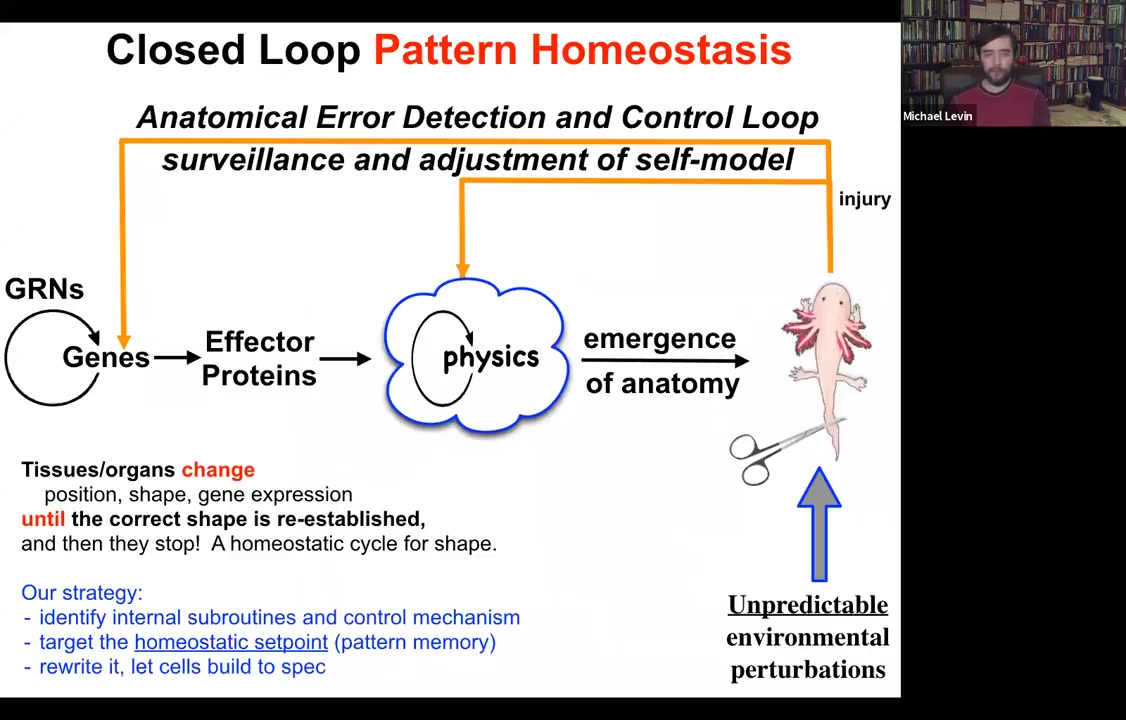
What we try to do is to come up with a scheme where there's more feedback here. When the system is deviated from its normal target morphology, be that with injury, teratogenic drugs, or pathogens, feedback loops kick in both at the level of genetics and physics, and we're going to talk about this physical one here, that try to minimize the error, the delta between where we are now and where we need to be. This is a classic homeostatic loop. It's what the thermostat in your house does. It measures against a set point, and if error is beyond a tolerable amount, it will undertake corrective action.
Several things to note here. The first is that feedbacks are not new in biology. There's something different here. The first is that the set point of this process: every homeostatic system has to have a set point towards which it tries to reduce error. The set point here is not a single number or a scalar like pH or a metabolic hunger rate. It's a complex set of information that in some rough, coarse-grained way describes what a correct anatomy should be.
The other thing is that this is very much a goal-directed process. When I say goal-directed, I don't mean something magical or mysterious. It's goal-directed in the cybernetic sense. We've had devices that are goal-directed agents since the 40s and 50s. All it means is that it's able to execute this error minimization loop.
But the cool thing about systems that work like this is the following. When you want to change what they do, you don't have to be rewiring the hardware back here. If you understood how the set point was encoded, you could change the set point and get the exact same cells to build something different. That's the amazing thing about any system that utilizes this kind of architecture, that you can make changes without even necessarily knowing how all the parts of the loop work. All you have to do is rewrite the set point.
So our strategy for some years now was this. We tried to identify some of the mechanisms at work here, in particular, to understand the homeostatic set point. How does the tissue store the pattern towards which it tries to remodel? Then let's rewrite that pattern and let's let the cells build. That's part of trying to decode this aspect of the collective behavior of these cells.
Let's talk about how we've been doing it. We have started looking at something called developmental bioelectricity. What I'm going to show you now are, first of all, the methods. How do we do this? and then show you some proof of principle applications.
Why is this interesting? I want to point out at the beginning that all cells in vivo sit within this complex morphogenetic field of information that tells them what to do as part of a larger unit.
Slide 19/54 · 18m:16s
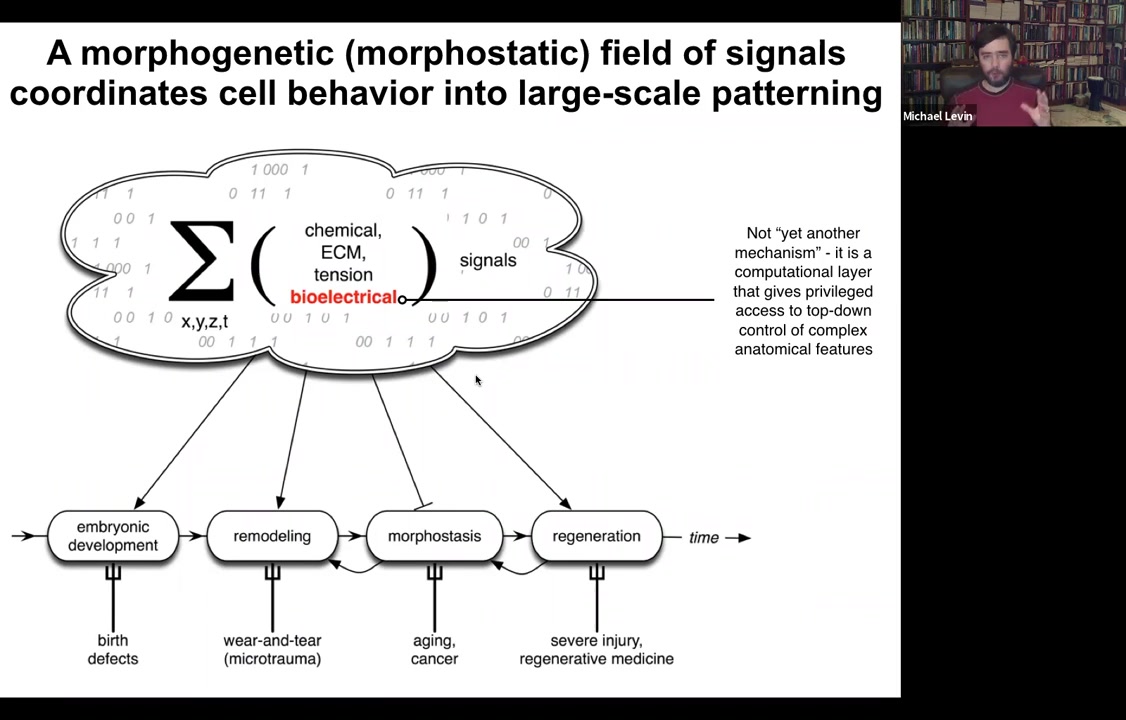
This information comes in many flavors. Chemical, extracellular matrix, tensions and stresses, biomechanics, all kinds of things, and bioelectricity.
Bioelectricity doesn't do this alone. It works with all of this other stuff. I'm going to focus today on bioelectricity because it has a particular aspect to it, which is that it's not just another piece of physics that you need to know to understand anatomies. It's actually a computational layer that gives some privileged access to the control of complex anatomical features. It is the medium in which the computations are being made to make decisions about the length and the size and the shape of things. Accessing this bioelectricity gives you a remarkable insight into what's going on beyond bottom-up molecular mechanisms.
Slide 20/54 · 19m:13s
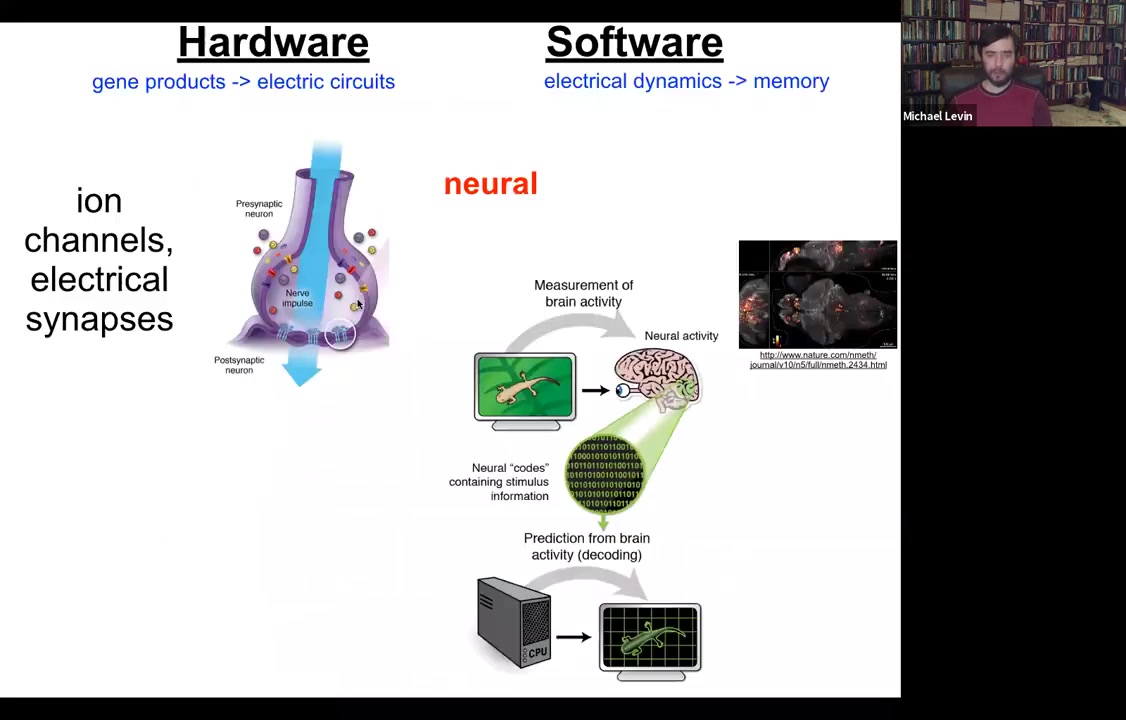
The kind of inspiration for how to think about this we took directly from neuroscience. In neuroscience, this story will be familiar to everybody. The hardware is simple. You have a collection of cells. Each cell has ion channels in the plasma membrane. They set voltage values across the membrane; that's the resting potential, which can go up and down, and it can propagate to the neighbors via these electrical synapses.
Those kinds of networks underlie this amazing software. Here you can see this physiology in a living zebrafish brain as the fish undergoes cognitive activity. You can see here the electrical activity. The commitment of neuroscience is that if we were able to decode this, if we understood how to read this information, through a computational approach we should be able to figure out what the animal is remembering, what it's thinking about, and what decisions it's making. The semantic and functional cognitive structures are to be read out from the electrical activity of the brain.
That's what we believe.
Slide 21/54 · 20m:27s
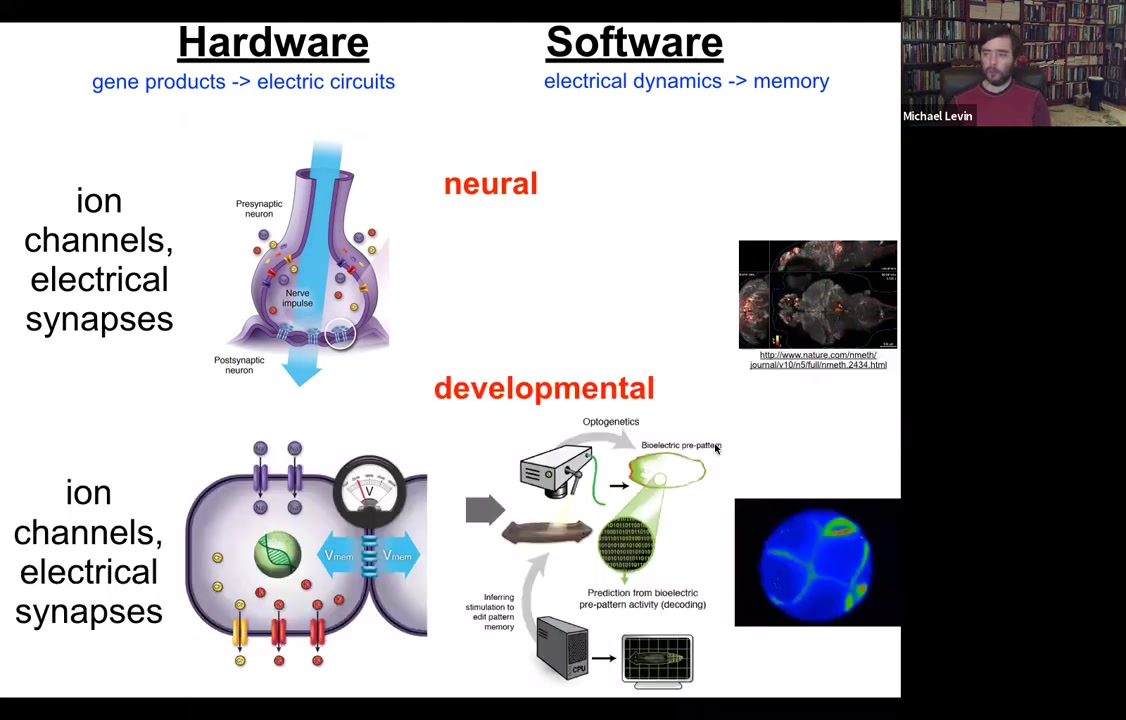
All cells do this. All cells have ion channels. Most cells have electrical synapses known as gap junctions to their neighbors. We have undertaken this project to try to do the same kind of decoding. Here's an embryo, and we would like to read all the electrical conversations that bind the individual cells into a collective that can undergo anatomical homeostasis towards large states, build a limb, build a kidney.
On the one hand, it's strange and surprising to many people to think about somatic cells as having this kind of neuroscience-like aspect to them, as if they're processing information, they're having goal states. On the other hand, if you ask where neurons came from, they didn't just appear out of nowhere. Evolution just sped up, optimized things that cells have been doing for a really long time, since around the time of bacterial biofilms. All of the components of neurons that are really important for this—the channels, the neurotransmitter machinery—existed before multicellularity, and even bacteria were using this to coordinate information across the biofilm.
Slide 22/54 · 21m:44s
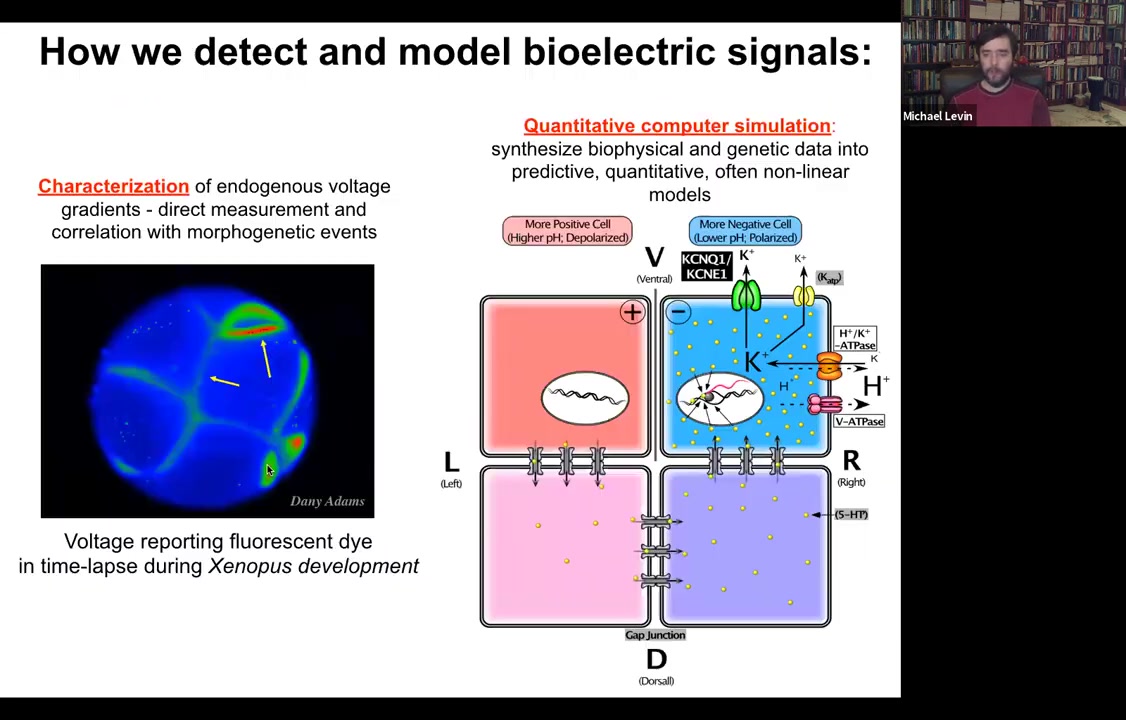
So we developed some tools to study this. The first is just how to detect and model them. This is a voltage-sensitive fluorescent dye, which reveals in the living state, non-invasively. You don't have to poke the cells with electrophysiological tools. Every cell now reports, around its membrane, all the voltages. This is an early frog embryo in time-lapse, sorting out which cells are going to be left and right, then dorsal, ventral, and so on.
In addition to that, we do a lot of computational modeling. If we know what ion channels and pumps are in the membrane, can we explain why the voltage is the way it is and why it changes as a function of time?
Slide 23/54 · 22m:28s
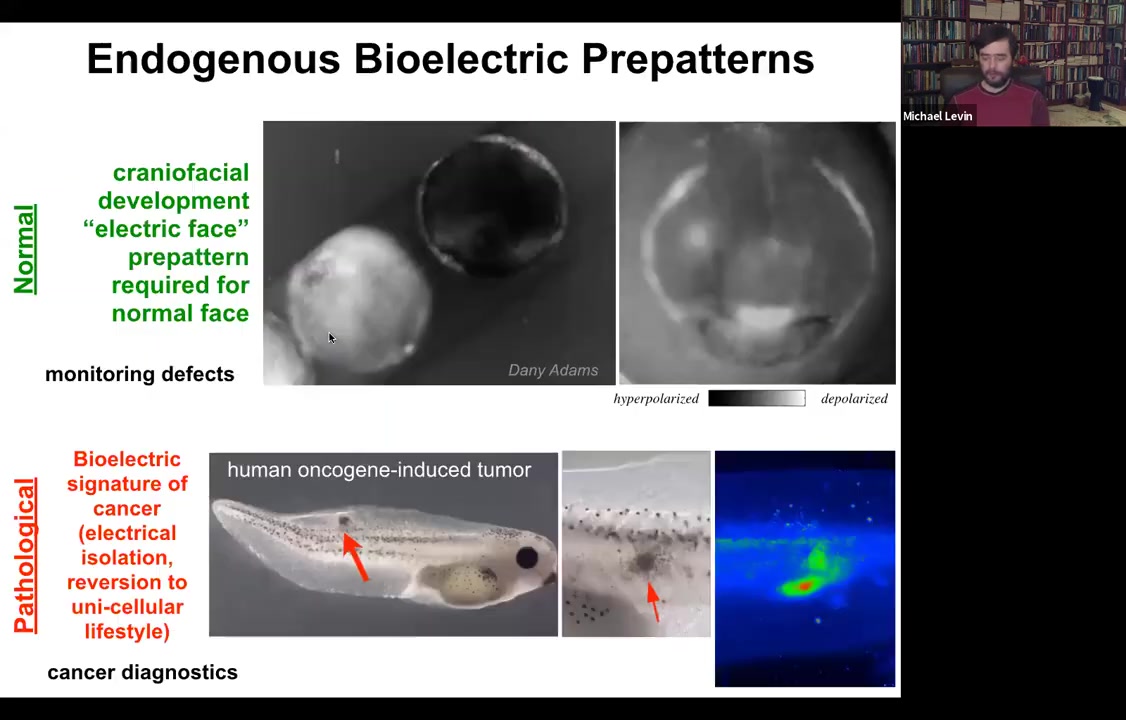
Let me show you a couple of native bioelectrical patterns. This is what we call the electric face. This was discovered by Danny Adams in my group. She used voltage dyes to look at the frog embryo putting its face together. This is, of course, a time lapse. This is one frame taken from that movie. Long before the genes come on to regionalize the face into the eye, the mouth, and everything else, all of the bioelectrical properties set up a pre-pattern, a scaffold that tells you where everything is going to be. Here's where the animal's right eye is going to be. The left eye will come in shortly. Here's the mouth. Here are some placodes.
This bioelectrical pattern is a native instructive pattern for gene expression and for anatomy. The reason we know it's instructive is because if you go in with optogenetics or other tools and change this electrical pattern, not move any cells, but just change the pattern, then you can get, for example, those Picasso tadpoles. You can move organs around at will.
This is a normal pattern that is necessary for correct craniofacial development. Here's a pathological pattern. This embryo was injected with a human oncogene, for example a KRAS mutation. Here it's going to form a tumor, and the tumor is going to start to spread. But before that happens, before it becomes histologically apparent, you can already see with this voltage dye that here are the cells that have depolarized and shut off the electrical connections to their neighbors in a way that's going to basically revert them back to their unicellular ancient lifestyle. Once you're electrically disconnected from this grid, the rest of the animal becomes just external environment, that computational boundary. Whereas before it was large, it was a group working on a liver or a kidney. It shrinks to the level of a single cell and all your goals become single-cell goals. You proliferate, migrate to where life is good. This ends up being metastasis and a conversion of the metastasis. This is a pathological pattern.
Slide 24/54 · 24m:38s
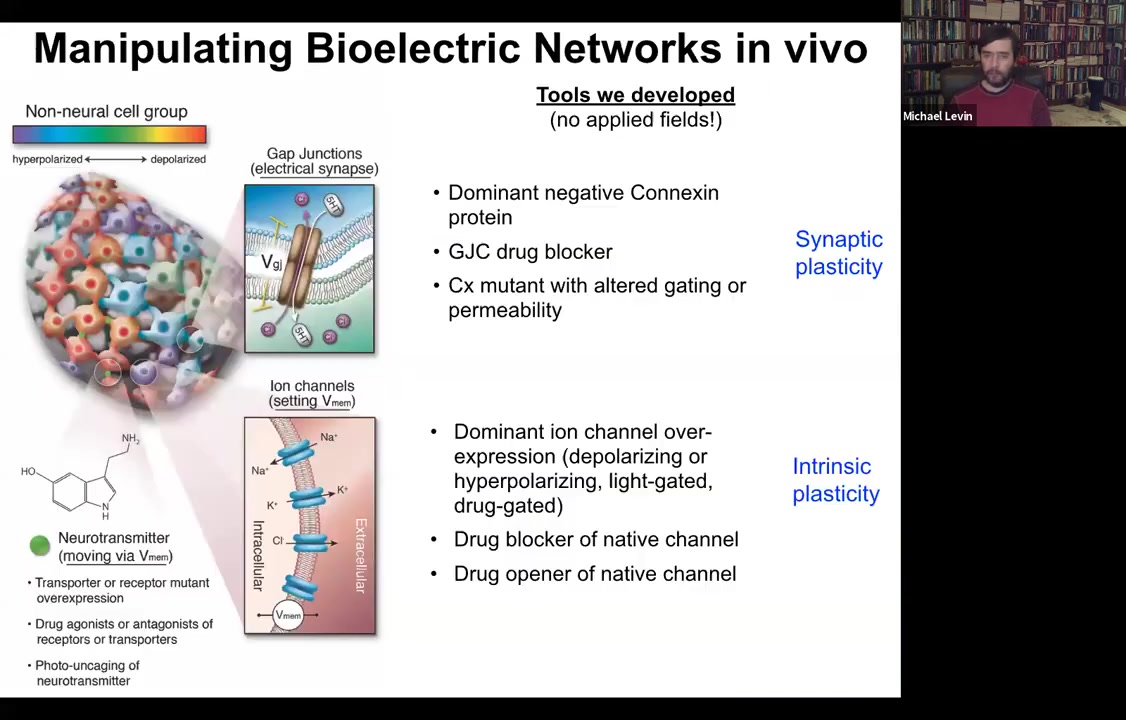
What we then developed was a set of tools that were designed to allow us to manipulate these bioelectrical gradients in vivo. If we thought they were functional, we ought to be able to control them to get the collective to do something different. It is important to say that we do not do any external field application. We don't have any electromagnetic components here. There are no electrodes. There are no waves. There's no magnetism. It's all molecular physiology.
The way it works is you have two things you can control in any tissue. You can control which cells electrically couple to which other cells. We target these gap junctions. We can mutate them. We can open them, close them, controlling synaptic plasticity, or we can control the various ion channels, again opening and closing them, to actually set the voltage of the individual cells. This would be the equivalent of some sort of intrinsic plasticity if this was neuroscience. We can use dominant mutations of channels, we can use drugs, we can use optogenetics, light, and so on.
This was back around the year 2000, when I was first setting up these tools and we wanted to control these bioelectric states, it was thought that resting potential is a housekeeping parameter. The standard assumption was that if we perturb it, the cells would die and you would get uninterpretable toxicity. That was the thought. I'm going to show you that, in fact, that's not at all what happens because these bioelectrical states are not just readouts and they're not just yet more molecular machinery, but they're actually the information-bearing medium for these large-scale modifications.
Slide 25/54 · 26m:30s
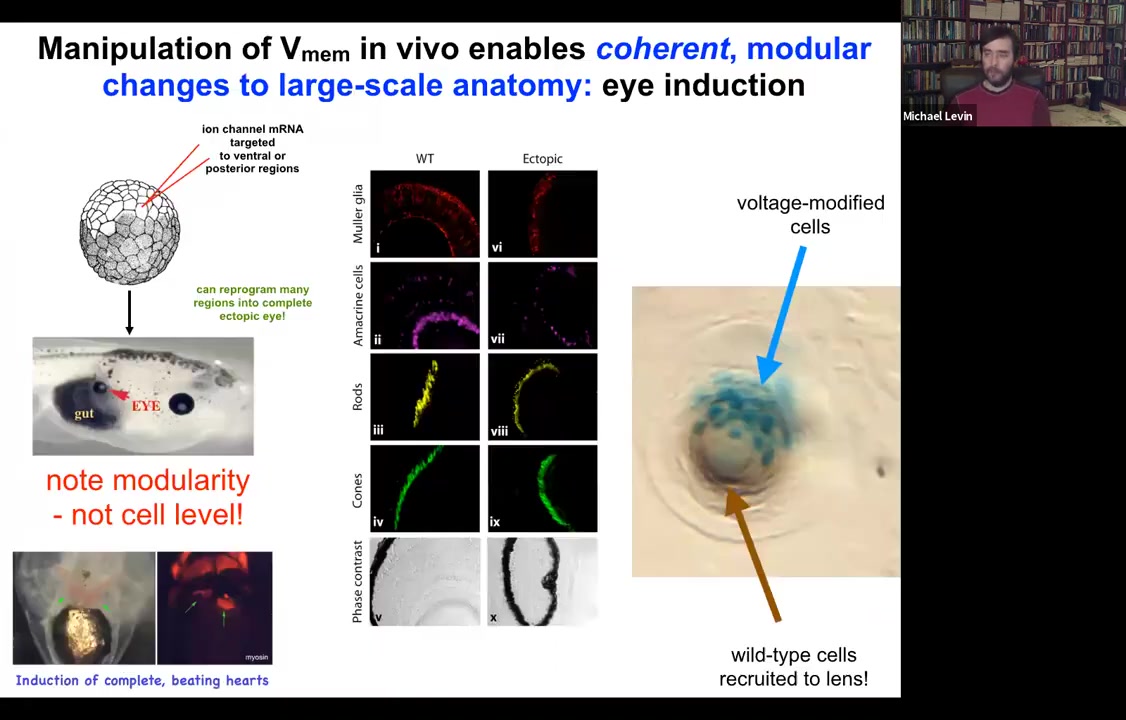
One thing you can do is you can take some cells in the frog that are going to become gut, so they're endodermal cells, and you can inject one of several types of ion channels that in this region that's going to be part of the gut will set up a voltage pattern, a distribution of voltage that's very similar to what happens when the native eyes come in. And sure enough, if you recapitulate that pattern somewhere else, the cells are using that pattern to decide what they're going to build. That pattern tells them to build an eye. They will build an eye out of endodermal cells. Now, in the textbook, it will say that only anterior ectoderm is competent to become eye up here.
Slide 26/54 · 27m:08s
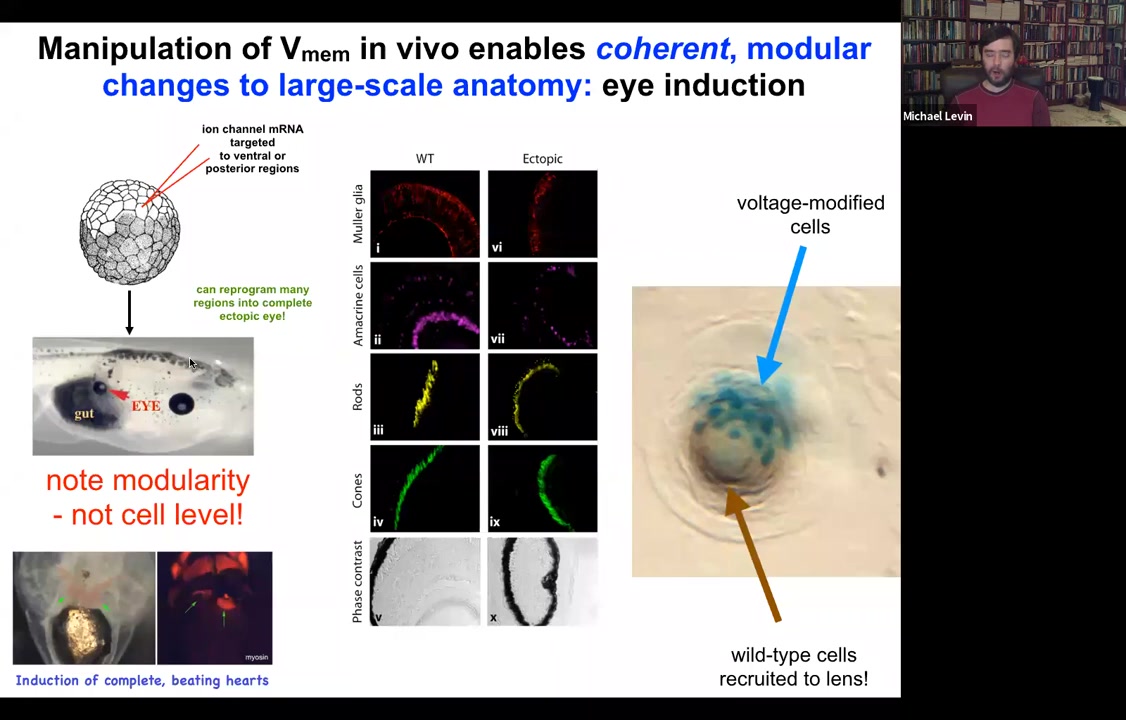
And that's true if you use canonical inducers like PAX6, the master regulator transcription factor. But if you go upstream of that and actually re-specify the bioelectric pattern, then you can induce these eyes anywhere, on the tail, in the gut, anywhere you want. And if you make these eyes, they can have all of the same components that normal eyes have. So they'll have lens, and retina, and optic nerve, and so on.
If you label the cells that you've directly injected with the ion channel, let's say we label them with beta-galactosidase, so they're blue. This is a cross-section through a lens sitting out in the flank somewhere. What you'll see is that there are two inductions here. The first induction is by us imposing a particular bioelectrical state that causes these cells to decide to become a lens. That's the first induction. The second induction is that these cells, with their aberrant voltage potential, actually recruit their neighbors, the brown cells here, which are not labeled, which were never directly modified by us, into this project of building this larger scale structure, this nice round lens. So it's a non-cell autonomous effect. And we see this again and again, that there's this ability of these bioelectrical signals to not just specify cell fate, but to actually specify organ type and position and things like that. So we can make eyes, we can make ectopic hearts. Here's a secondary heart.
We can make limbs and brains and some other things, and then many things that we can't yet make.
Slide 27/54 · 28m:44s
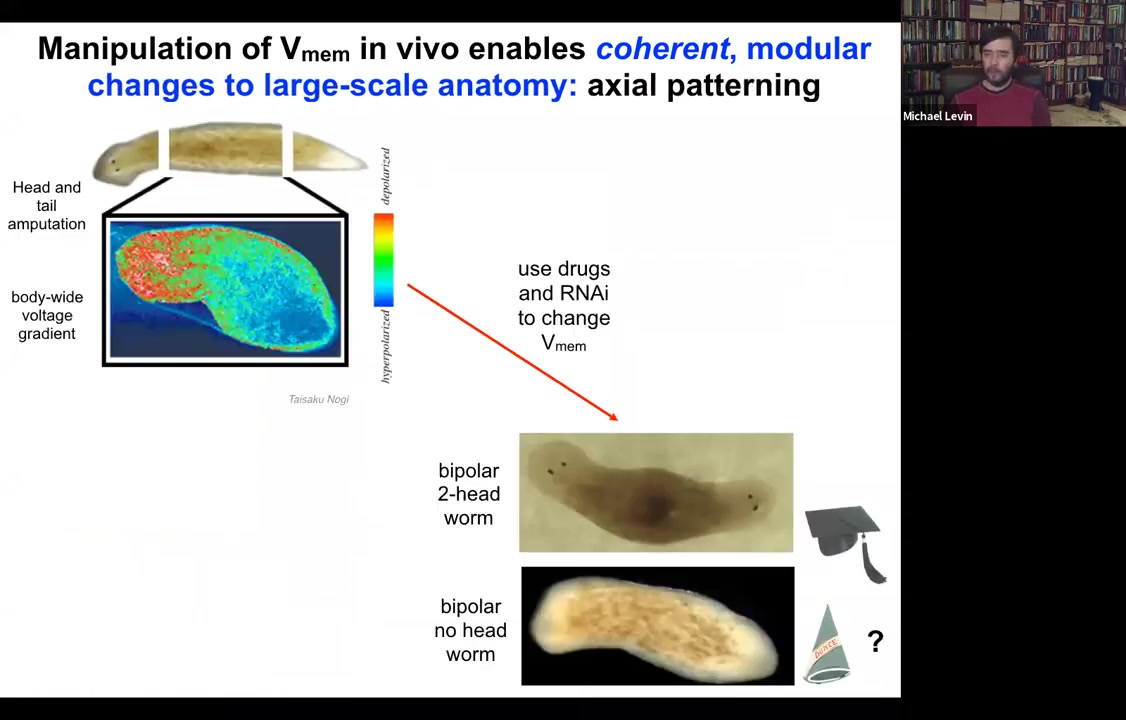
Let's look at another example. Here's a planarian. One of the most interesting things is that if you chop off the head and the tail, this middle fragment knows exactly how many heads it's supposed to have. It puts one head here, it puts a tail here. The way it does that is because as soon as you amputate, there's this voltage gradient that's set up where the red, the depolarized region tells the worm how many heads it's supposed to have.
Slide 28/54 · 29m:06s
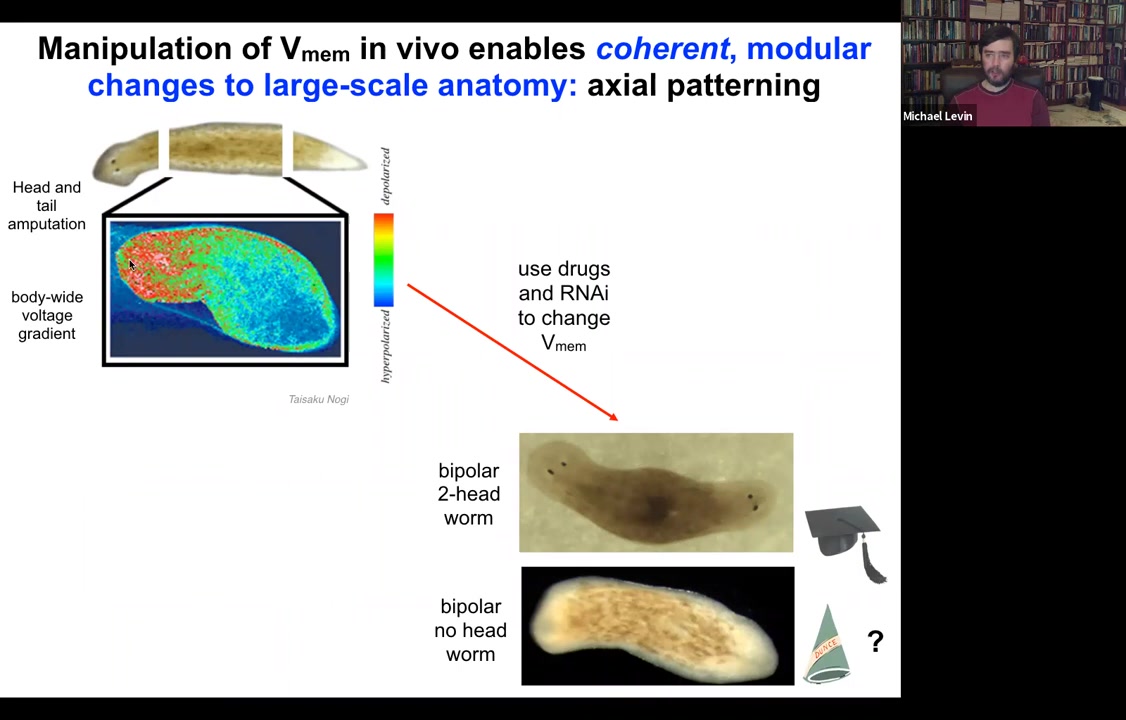
What we can do is we can go in and target this region using drugs, ion channel modifying drugs to depolarize this region.
When you do that, you get a two-headed animal. These cells are perfectly happy to make a head. The information of how many heads you're supposed to have comes from this electrical gradient. If you change the gradient, they'll happily make two heads. You can make two-headed animals. You can make no-headed animals.
What's important here is that there's no genomic editing here, so this is purely physiological, and I can ask this animal to make a second head of the correct type.
Slide 29/54 · 29m:52s
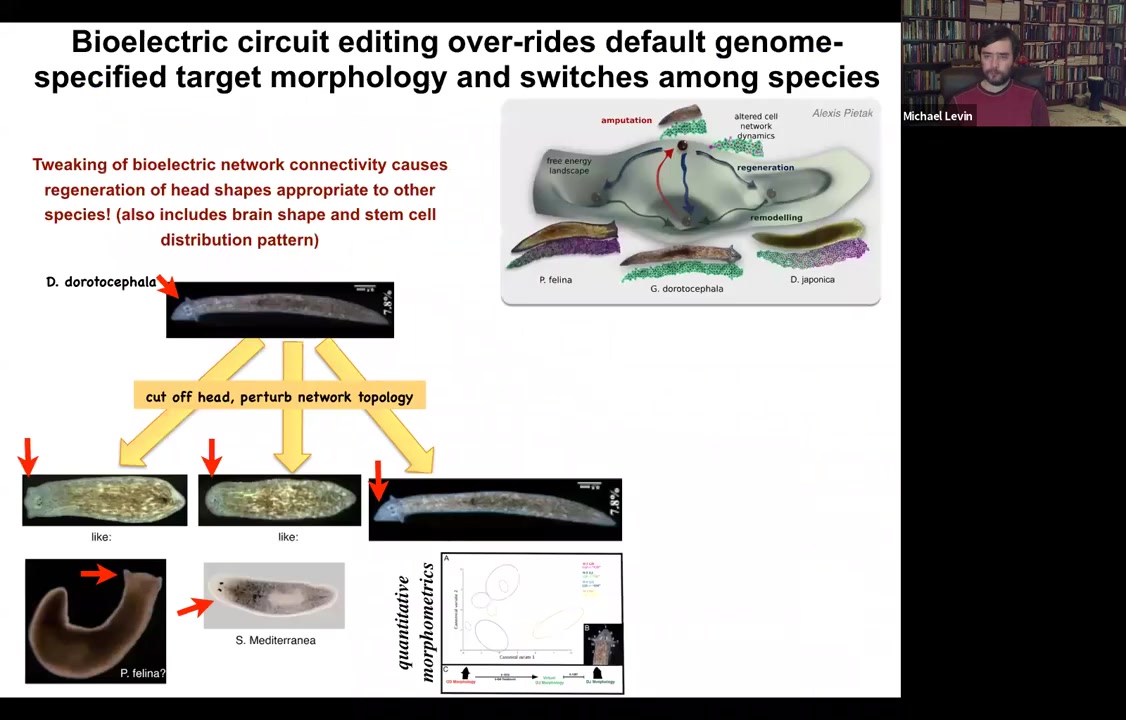
I can actually ask it to make heads belonging to other species. We started off with a triangular-headed worm like this, amputated the head and perturbed for about 48 hours the ability of cells to talk to each other electrically. What happens is the electric circuit is unable to settle to the correct attractor. When they do settle stochastically, sometimes they settle to the correct shape, but sometimes they make round heads or flat heads. In doing so, they recapitulate the state of flat-headed species like this P. falina or round-headed species like this S. mediterranea. It's not just the shape of the head, it's also the shape of the brain and the distribution of stem cells that is exactly like these other species. No genomic editing here, perfectly wild-type genome. These animals, once the electric circuit settles, there's a good chance that it'll settle into an inappropriate attractor that's actually associated with a completely different head shape that other species are using natively. You can also knock the system into attractors that are not being used by any system.
Slide 30/54 · 30m:58s

And you can really get far from the normal planarian shape. You can make animals that are spiky. You can make animals that are cylindrical, or a combination of a flat animal with a tube growing out into the third dimension. I'm going to show you in a minute that what this is all about is predictive control over large-scale anatomy. This is not about making weird, aberrant teratologies. The idea is that we can show how much of this is under bioelectrical control and then use computational techniques to gain good predictive control over it.
This is a good time to think about the role of the genome in the anatomy of this process. The typical metaphor that we hear is that the DNA or the genome is the software and that the cell is the hardware that interprets this.
That's not a bad analogy at the single cell level when you're thinking about phenotypes of proteins and pathways.
But at the level of anatomy, I would suggest a different metaphor. The different metaphor that works better to understand anatomical control is this.
Slide 31/54 · 32m:14s
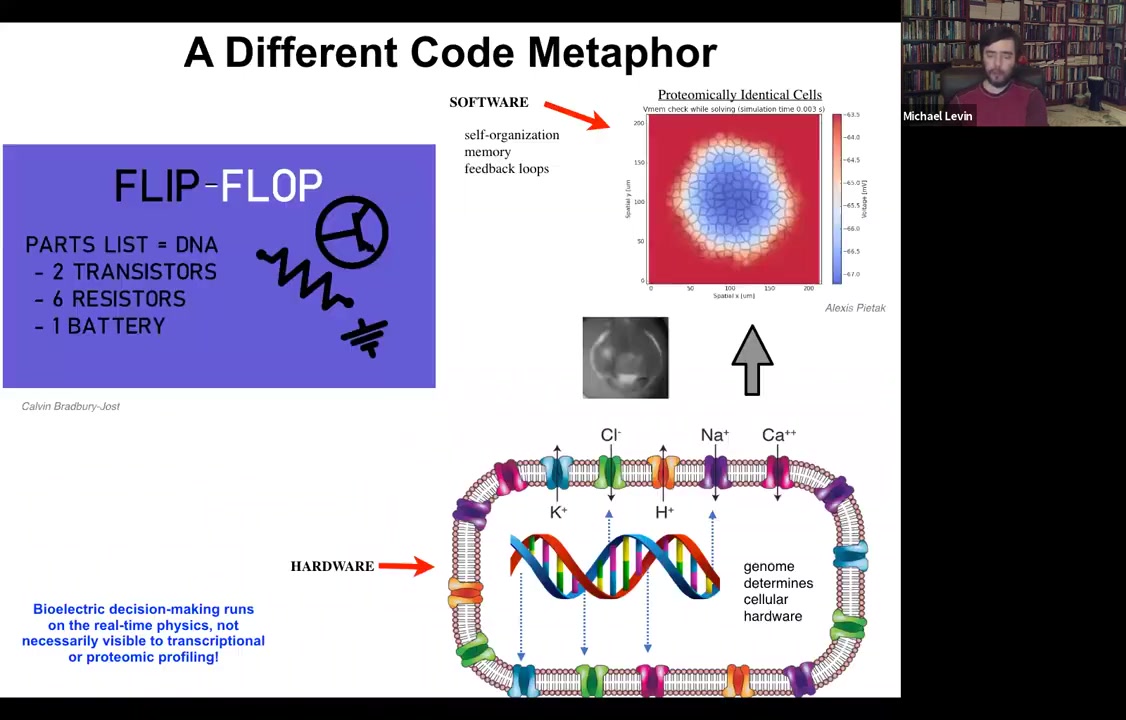
What the genome does is specify all of the hardware that the cell gets to have. All of the specific ion channels that are here are determined by gene expression. But we know from basic neuroscience and also from computer technology that if you make a very simple circuit, this is a flip-flop circuit, if you nail down the hardware, you can have a circuit that can store multiple different pieces of information depending on the current flow through the system and store it stably, which is a memory. A flip-flop is a basic kind of memory. And you don't need to move or change any of the hardware in order to store a zero or a one in this very simple electrical circuit.
So in fact, with our modeling, you can see that the same thing is true if you set up a field of cells where every single cell is expressing exactly the same set of ion channels. What you actually make is an excitable tissue where very rich patterning can take place. It's a little like Turing patterns. There's spontaneous symmetry breaking and self-organization. But all of this happens with a completely constant proteome. You don't need to change these ion channels because they can open and close. And now all of the dynamics are at the electrical circuit level, not with the proteins underneath. The prediction of this way of thinking about it is this: you should be able to edit the software, meaning change the information stored in the system while keeping the hardware constant.
In particular, the piece of information that we want to change is the set point. We want to change the set point towards which cells are working in this error minimization scheme.
Slide 32/54 · 34m:03s
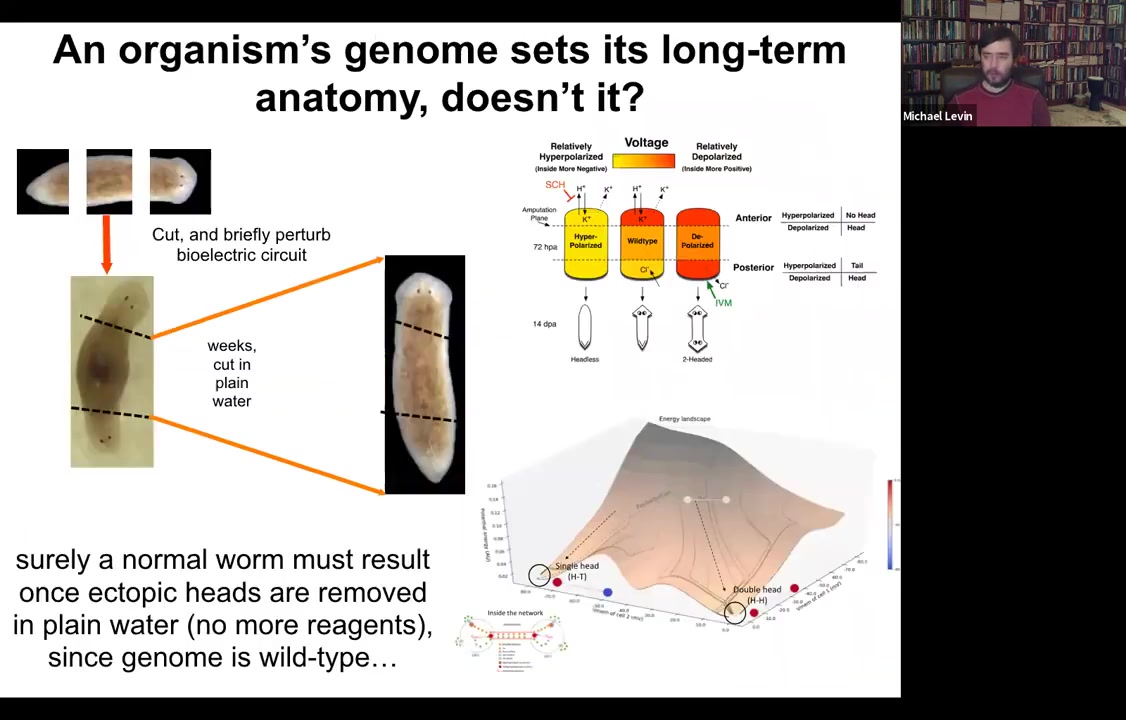
Let's think about how to do that. Here's our normal planarian. We've cut off the head and the tail. We got the middle fragment. We've perturbed the electric circuit according to this model that we've developed. Here's your two-headed animal.
We ask a simple question. Here's our two-headed animal. We give it a couple of weeks to get everything settled down. Cut off the primary head. Cut off this crazy ectopic secondary head. Some people think you somehow epigenetically reprogrammed this posterior tissue. Fine, we'll cut it off. We'll throw it away. All that remains is a normal middle fragment here that didn't have any head tissue in it. We do this in plain water, no more manipulations of any kind. The genome is wild type. We haven't edited the genomic sequence at all.
Surely the prediction would be that you should get back to a single-headed worm. Once you cut off this thing, you should be back to a single-headed worm. The interesting thing is that if you model the state space of the electric circuit that's involved here, what you find is that there are multiple stable points. One is here, a very stable point around the single-headed shape, but there's another stable point around the double-headed shape. That suggests an interesting idea: is it possible that when you amputate this thing, the bioelectrical circuit will still remember that it needs to make two heads.
Slide 33/54 · 35m:22s
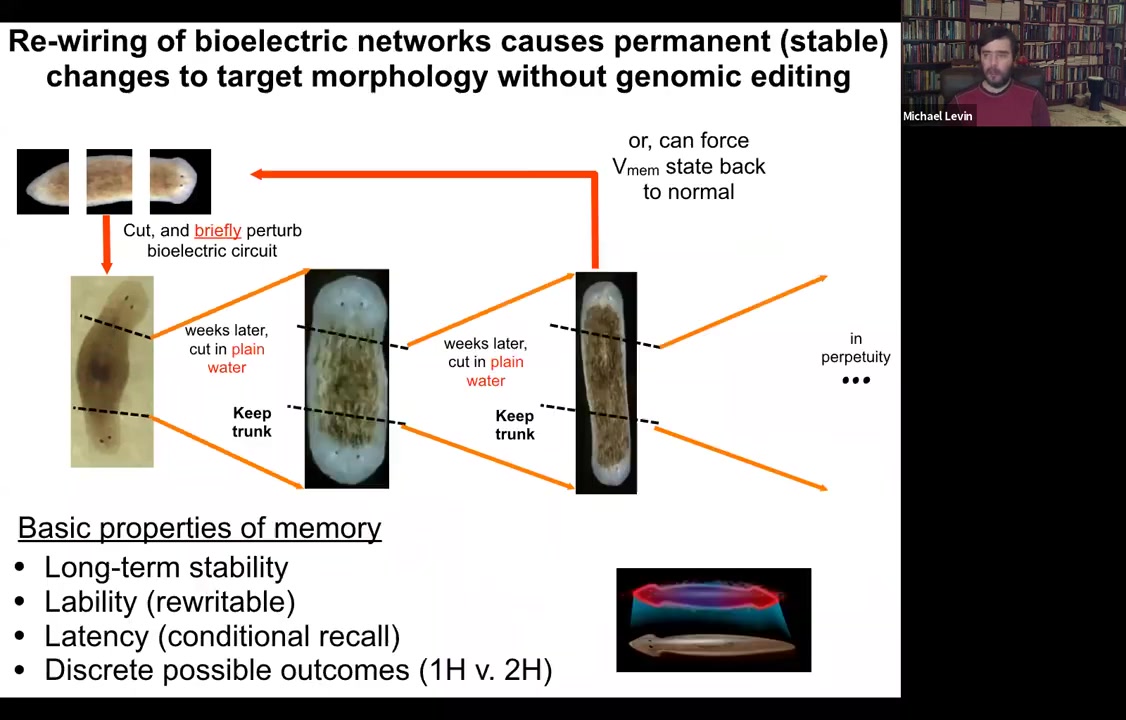
And sure enough, that's exactly what happens. When you amputate these in perpetuity, as far as we can tell, forever, two-headed animals will continue to give rise to two-headed animals, even despite their wild-type genome in plain water, no more manipulation. And we can, in fact, if we want to, we can set the bioelectric circuit back to normal, and it goes back to being a single-headed animal. So a normal planarian body, as you're going to see, is able to store a couple of different, at least, probably more, but we've nailed down two, ideas of what a correct planarian is supposed to look like.
Slide 34/54 · 36m:00s
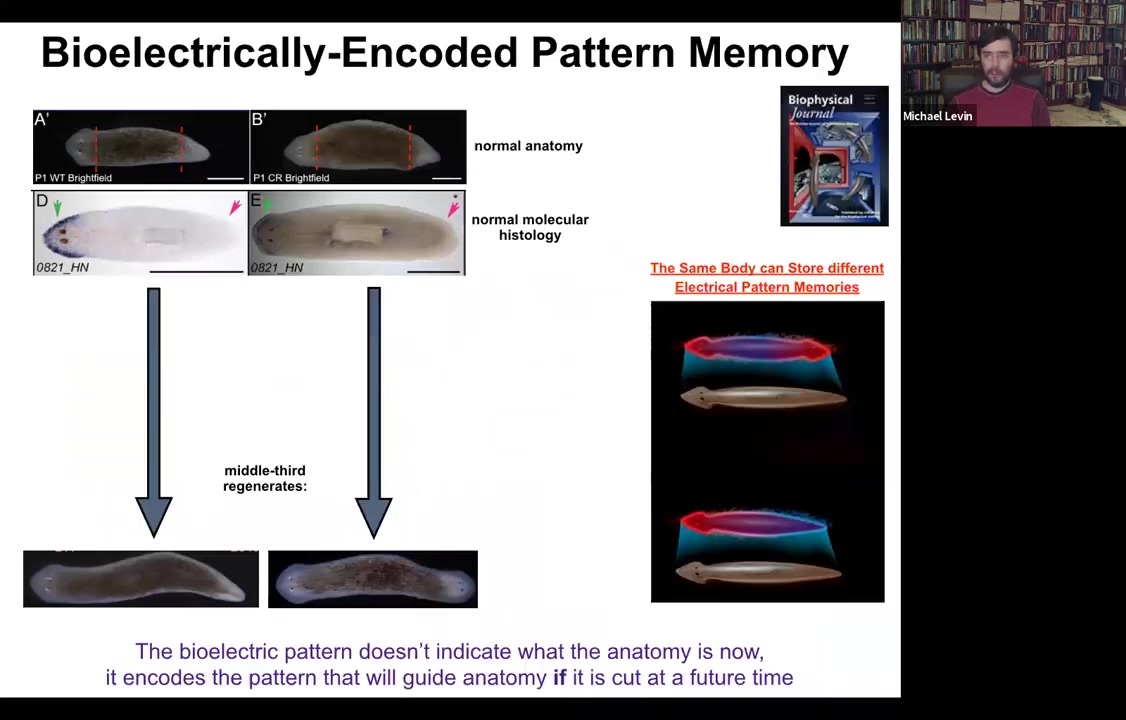
Here's a single-headed planarian. If we look at the molecular markers, the anterior marker is here in the head, not in the tail. If you amputate, you get a perfectly normal single-headed animal. Here's another anatomically normal animal.
Slide 35/54 · 36m:16s
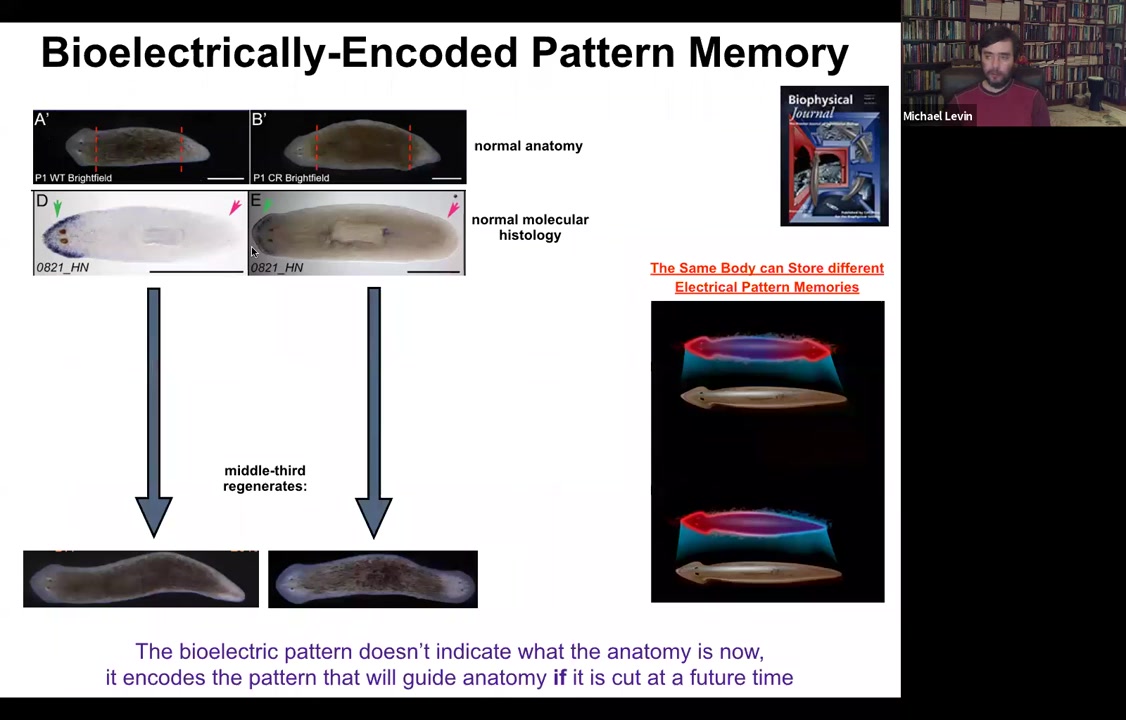
Again, molecular markers are all normal, anterior in the head, no anterior marker back here. But if you amputate this one, suddenly you get a two-headed animal. Why would you get a two-headed animal? Because in the meantime, what we've done is we've bioelectrically changed the stable pattern to say, no, now you need to have two heads. Now, here's the critical part of this: the electrical map here is not a map of this two-headed animal. This electrical map is a map of this one-headed animal, meaning that the bioelectricity distribution is not simply a map of whatever the anatomy is doing. The same single-headed anatomy can store at least one of two different stable representations of what it's going to do if it gets injured in the future. If you're interested in neuroscience and how counterfactuals are remembered in the brain, this is the evolutionary precursor to being able to remember things that are not happening now, either things that happened before or might happen in the future. It's a counterfactual memory. It's the ability to store a representation of a worm towards which the fragment is going to build if it gets injured. So it sits perfectly comfortably as a single-headed animal with this pattern until you cut. At that point, that memory is no longer latent and it actually becomes functional.
And that's how we get this propagation of these two-headed worms.
Slide 36/54 · 37m:43s
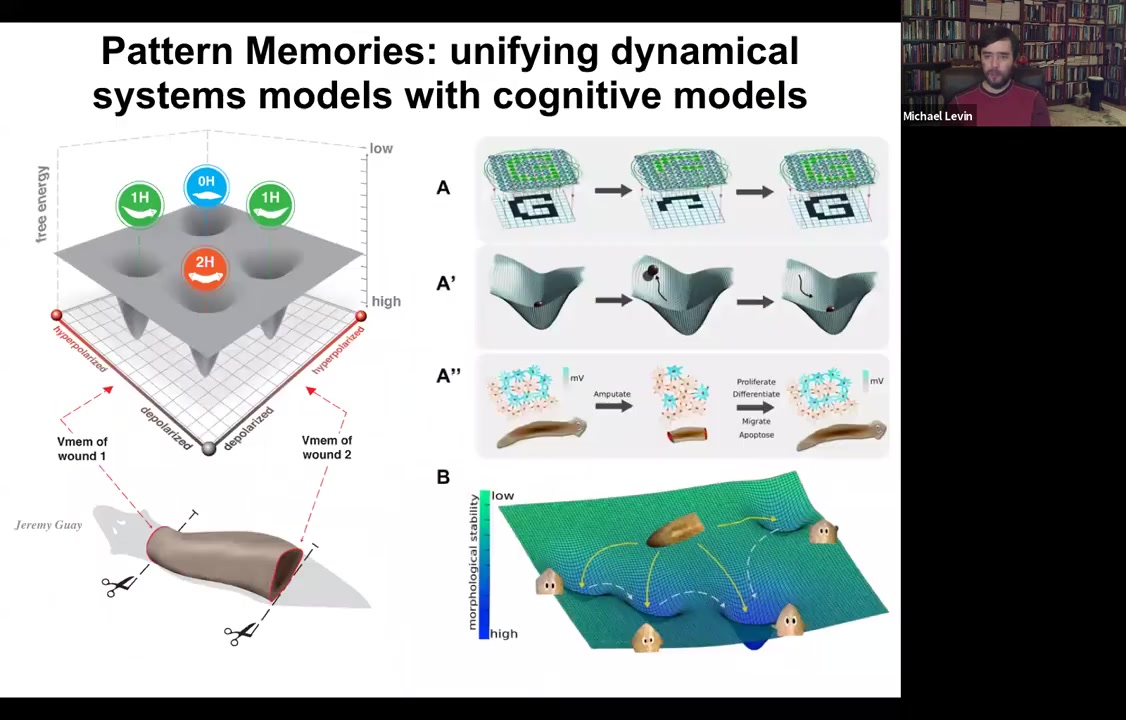
So we are building very quantitative molecular biophysical models to understand the state space of the circuit, to understand how the circuit minimizes free energy and eventually lands in one of these attractors, hopefully the right one, but not necessarily. And tying it into exactly the same formalism for how people in machine learning think about networks, electrical networks that can store patterns and can repair those patterns when pieces of information are missing. This is all very well-trodden ground in machine learning. And we think that some of these same strategies that we use now were discovered by evolution long ago.
And beyond worms, this is the beginning of our roadmap for regenerative medicine.
Slide 37/54 · 38m:39s
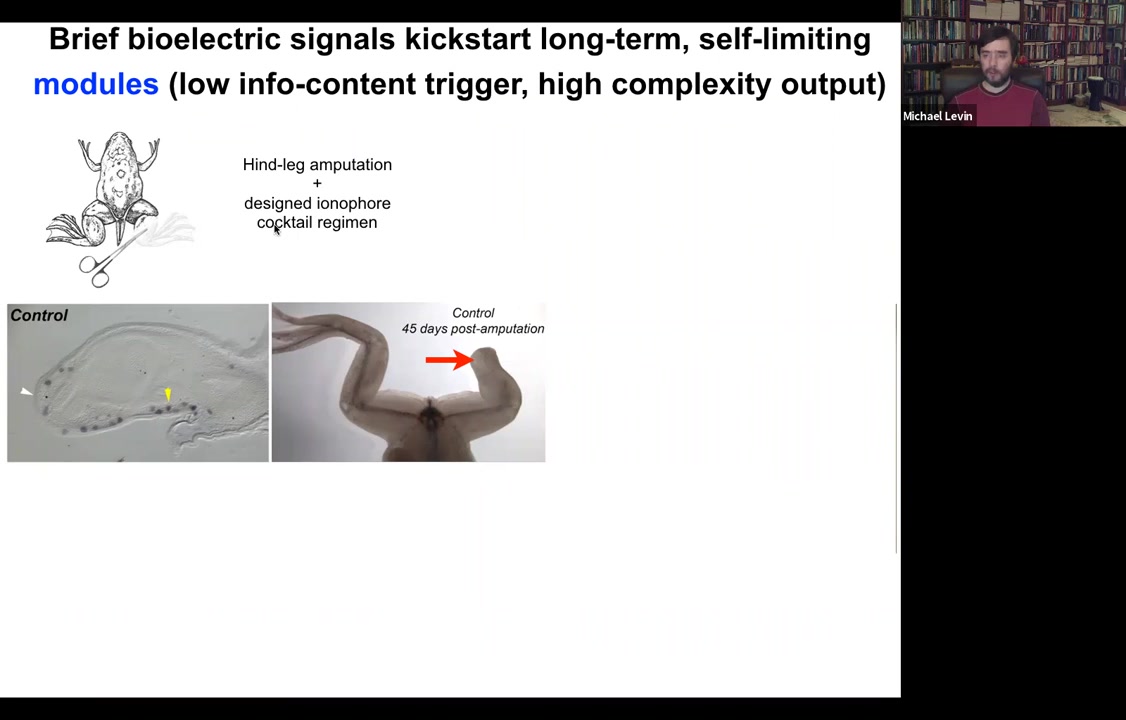

Because what we found is that, for example, if you have a frog and the frogs do not regenerate their legs the way that salamanders do, you amputate the leg, 45 days later, there's basically nothing. There's no blastema and there's no regenerate. What we can do is we can apply a cocktail of ion channel drugs that serve as a stimulus to kickstart a very complex cascade.
Slide 38/54 · 39m:02s
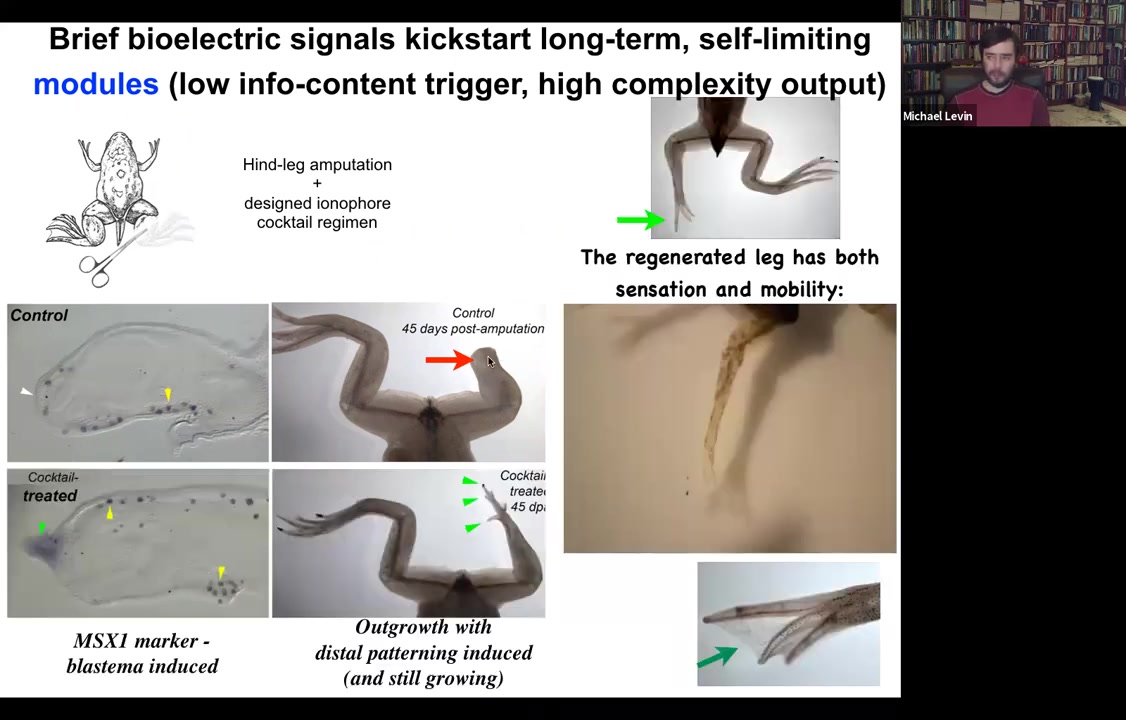
What you see here is that immediately the MSX1 blastema marker becomes turned on, and the leg starts growing. Already you have the toes and here's a toenail. And eventually you get a very respectable leg here and it's touch sensitive and motile. The animal can use it.
There are a couple of interesting things about this. One is that much like when we created an eye or induced a second head on a planarian, inducing this leg, we don't provide all the information to micromanage the process. We have no idea how to build a planarian head from scratch or how to make an eye or how to make a leg. What we have found is a trigger for a subroutine that this tissue already knows how to do. And the decision for what it's going to do, scar or produce various types of organs, is part of an electric circuit that we can guide.
The second thing is that, in keeping with this idea from the beginning that one of the hallmarks of the collective intelligence of these cells in responding to novel situations is that they can get to the final outcome through novel paths. If you actually look at this intermediate stage of this leg, this is nothing like what frog legs look like when they're developing in the first place. It eventually gets to a very good frog leg shape here, basically indistinguishable from the control. But the intermediate path through morphospace, the ability of the system to navigate that anatomical morphospace is amazing because it does not follow the same path that frog limbs normally follow where they make a paddle and then there's apoptosis that kills off the space between the digits. It's not what they do. They grow; it's almost like a plant meristem where you get the central stalk with this nail and then off to the side you get these toes and eventually the whole thing remodels into a leg. The remarkable ability to get its job done in a different way.
Slide 39/54 · 41m:01s

This is not just things that apply to frog and worm. We've done some work with David Kaplan and Lauren Black and others on human mesenchymal stem cells and cardiomyocytes. You can control the kinds of things that you can see in vitro, differentiation, proliferation, wound healing. But I think that's not what the bioelectricity is for. What the bioelectricity is really for in terms of why evolution uses it and why we should be interested in it is because it allows us to exploit the modular nature of the anatomical decision-making, because we can control very large-scale outcomes, not micromanage the details.
Here I have to do a disclosure. David Kaplan and I are both co-founders of this company called Morphoseuticals Inc. where we're trying to take the things that we learned about frog leg regeneration and move them towards mammals. Our goal is we're now in rodents trying for limb regeneration using the same strategies and these wearable bioreactors. The idea is to kickstart regeneration, not micromanage it, not try to babysit all of the different cell types and growth factors, but kickstart a trigger very early on and convince the cells with an aqueous protective environment that regeneration can proceed in the way that it would in a salamander or in an embryonic mammal during this adult phase. That's where one of the research programs is going.
Slide 40/54 · 42m:38s
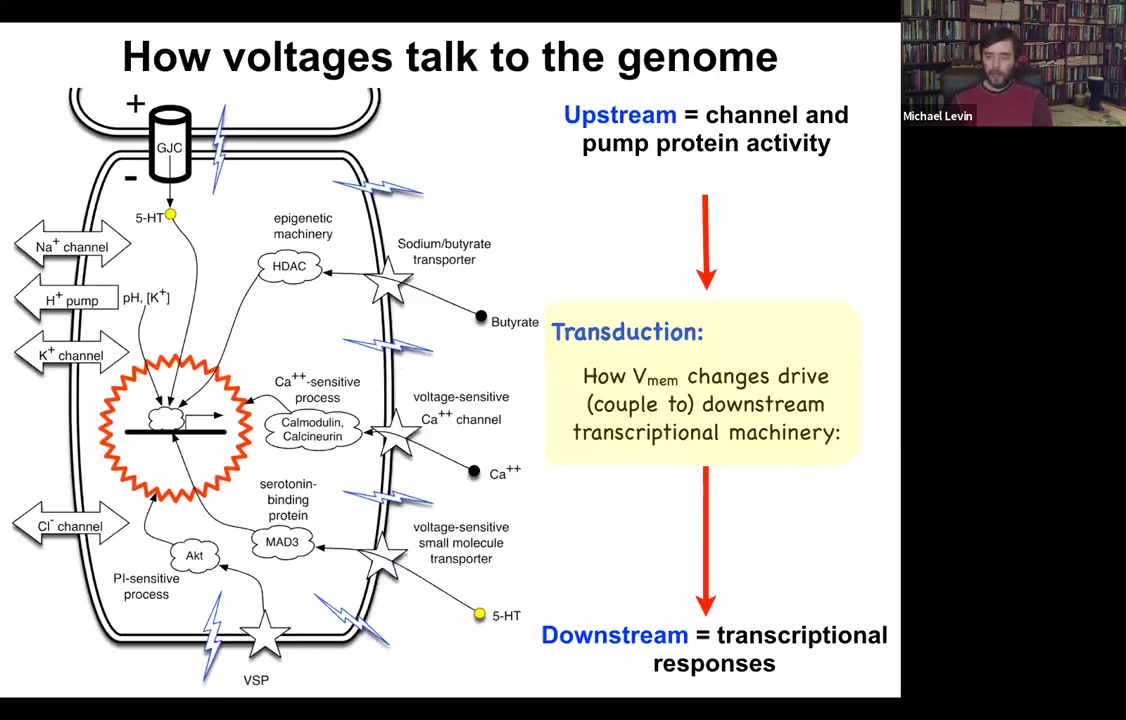
I should point out that it is already known how voltage change impacts the transcriptional machinery. People often say that it would be great if we understood how voltage controls gene expression. We do. We already know about six ways that voltage change transduces down into second messenger pathways and controls gene expression. These include typical mechanisms like calcium, which neuroscience is very familiar with, other things like voltage-sensitive phosphatases, and neurotransmitter transporters. We already know how this works at a single cell level. This transduction machinery has been dissected.
Slide 41/54 · 43m:23s
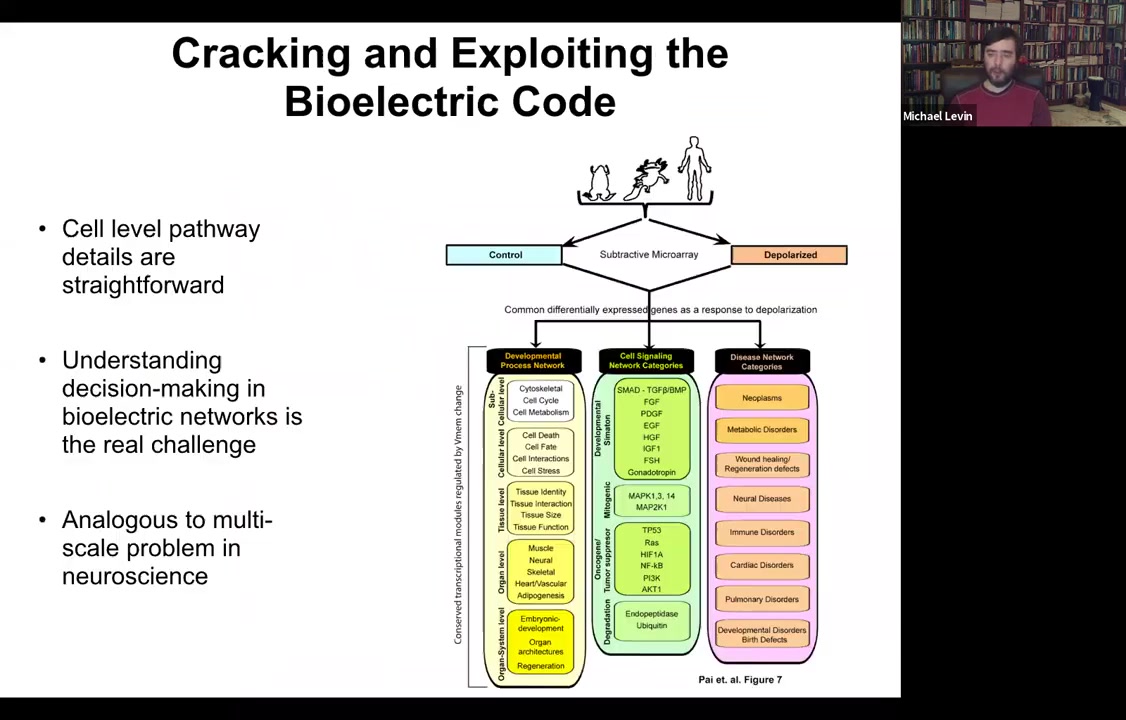
Also some of the downstream targets have been identified. The genes that are controlled by this — all of your favorite BMPs, Sonic Hedgehog, FGFs — are in fact the redistribution of morphogens in the examples that I showed you. All of this is known both from specific candidates and through unbiased RNA-seq and microarray experiments.
We know this, but what's interesting is that these answers about how it works at a single-cell level have been fairly unhelpful in understanding the large-scale picture that we're really interested in. It's analogous to this multi-scale problem in neuroscience. You can track the pathway, and in every paper you have to go from the channel to the transduction machinery, which genes are downstream. But that actually leaves open some much deeper questions about how the collective makes decisions.
Slide 42/54 · 44m:17s
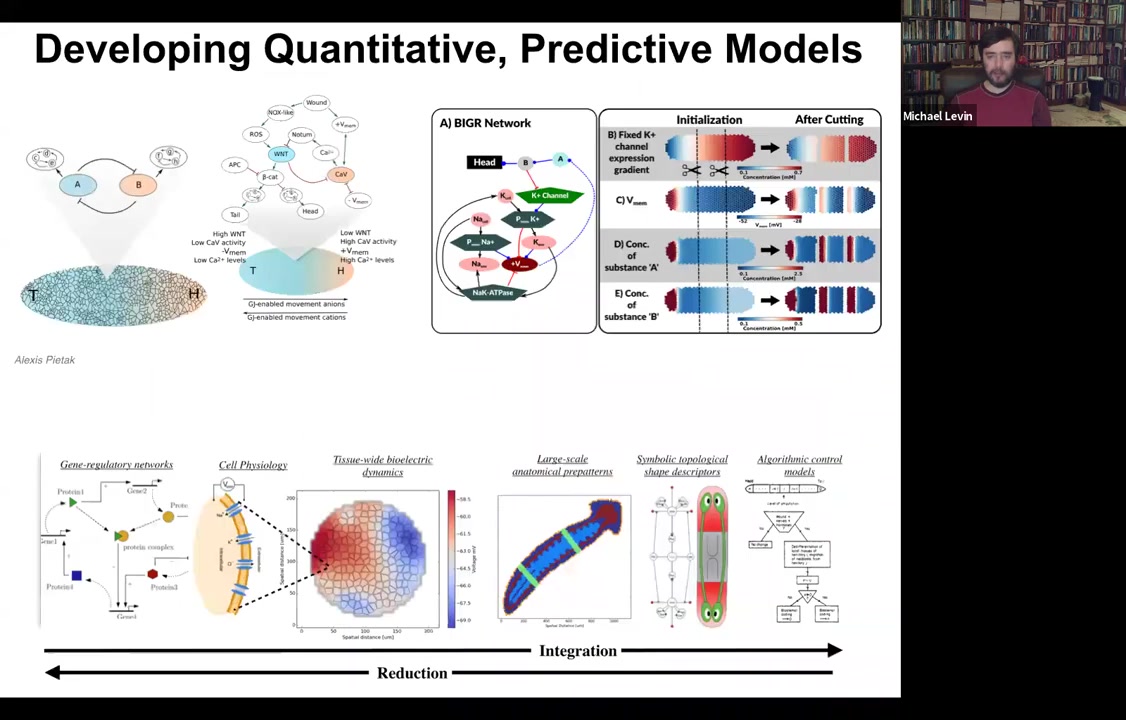
We've made the simulator, and this was created by our center member and collaborator, Alexis Pytak. What it allows us to do is to build models of both molecular networks like GRNs together with the various biophysics steps that happen. You can simulate every cell having these various circuits. Then you can try to build a full stack model that integrates from the molecular steps to the tissue level physiology. Every cell has various channels. What is the electrical pattern going to be in this tissue? What does that look like on an axial polarity level? Where's the head, where's the tail?
Can we extract from this the actual rules of the decision making? Can we turn this into a kind of algorithmic description that looks like this that makes it much easier for us to control the process? Because now this is almost like a high level programming language where we can see here are the measurables and here are the decision points that the circuit is using to control large-scale patterning modules like make a head, make an eye.
What we're doing is literally integrating every part of this so that you can build it up and then exert all your manipulations at this level. You could have your simulator and ask questions: why does the head-tail gradient rescale when you cut the planarian into four pieces? How does that work? You can't see that by drilling down into single cells. You have to come back out with that information and build these collective kinds of models.
Slide 43/54 · 45m:52s
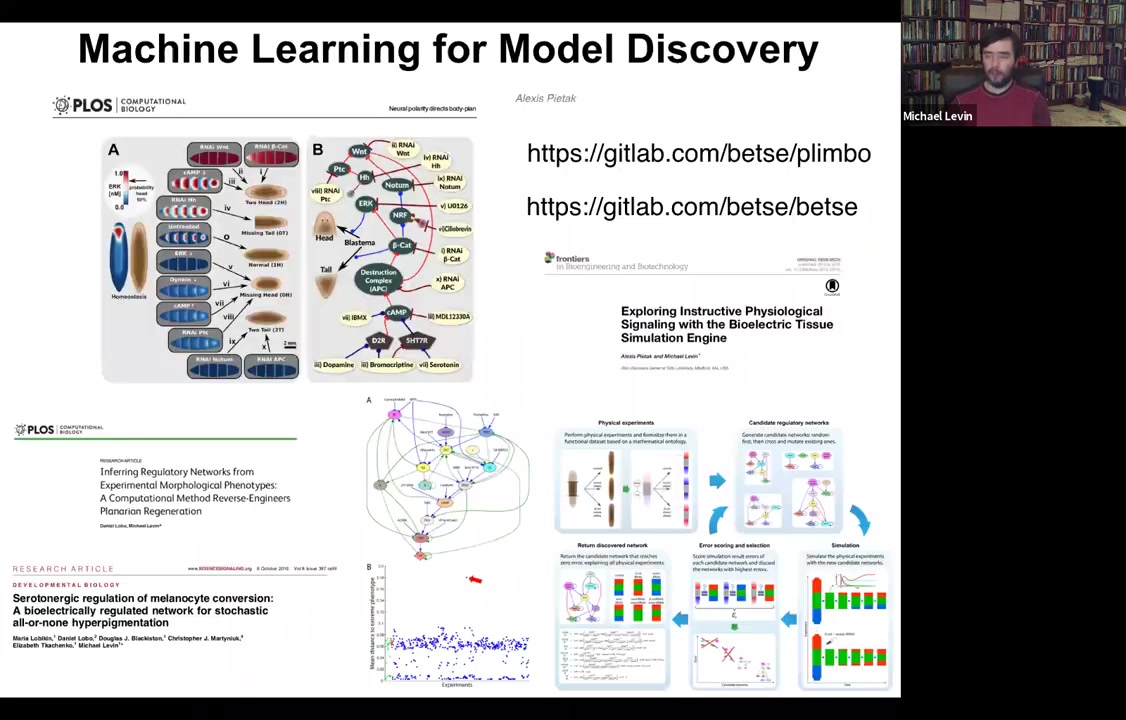
And the other nice thing about these models is that they are the perfect fodder for machine learning approaches to do two things: to infer better models from data. And we've done some of that. And then interrogate those models for interventions. And so here's some software that anyone can play with if you want to download and simulate all this. And we've shown that you can use machine learning to find models that are really good at explaining functional data sets. In other words, you did something to the system and then something else happened. What is a model that explains that? And then find these needle-in-the-haystack interventions that are ways in which you could perturb it to get it to do what you want. So that is the future of using this for regenerative medicine.
So let's talk specifically about a couple of applications in the last few minutes.
Slide 44/54 · 46m:49s
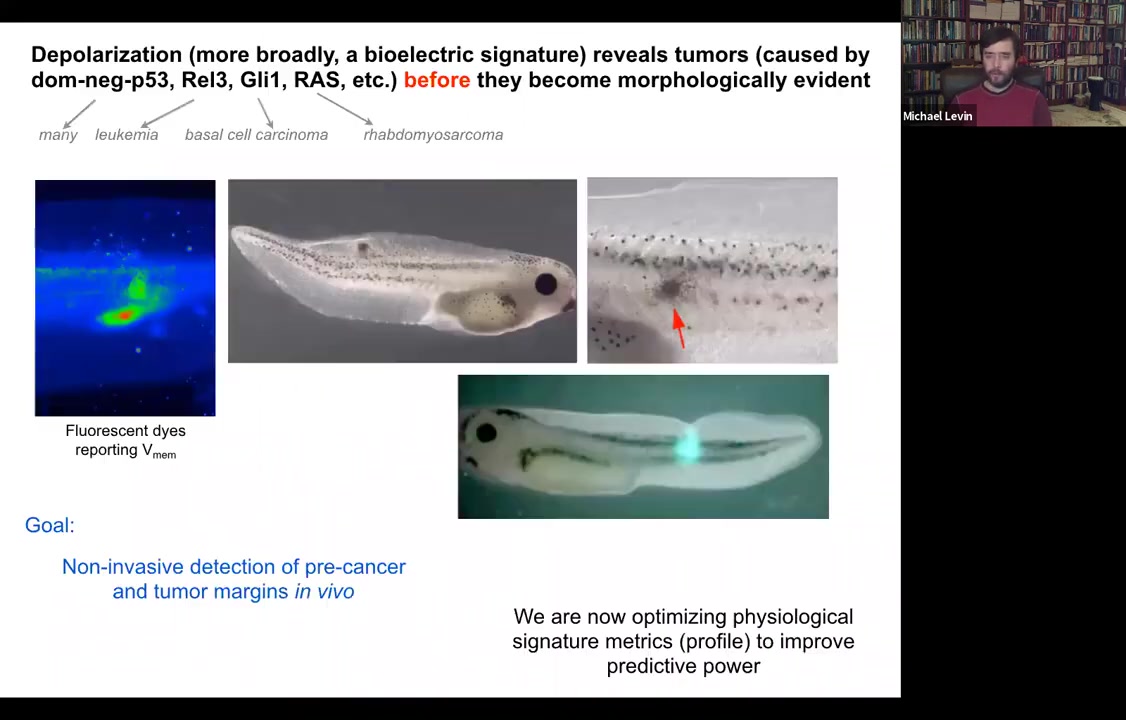
The cancer issue. I've shown you that you can track the shrinking of these kinds of computational boundaries from the organ or whole-body scale to single cells. You can use this as a diagnostic modality, screening for cancer.
Slide 45/54 · 47m:17s
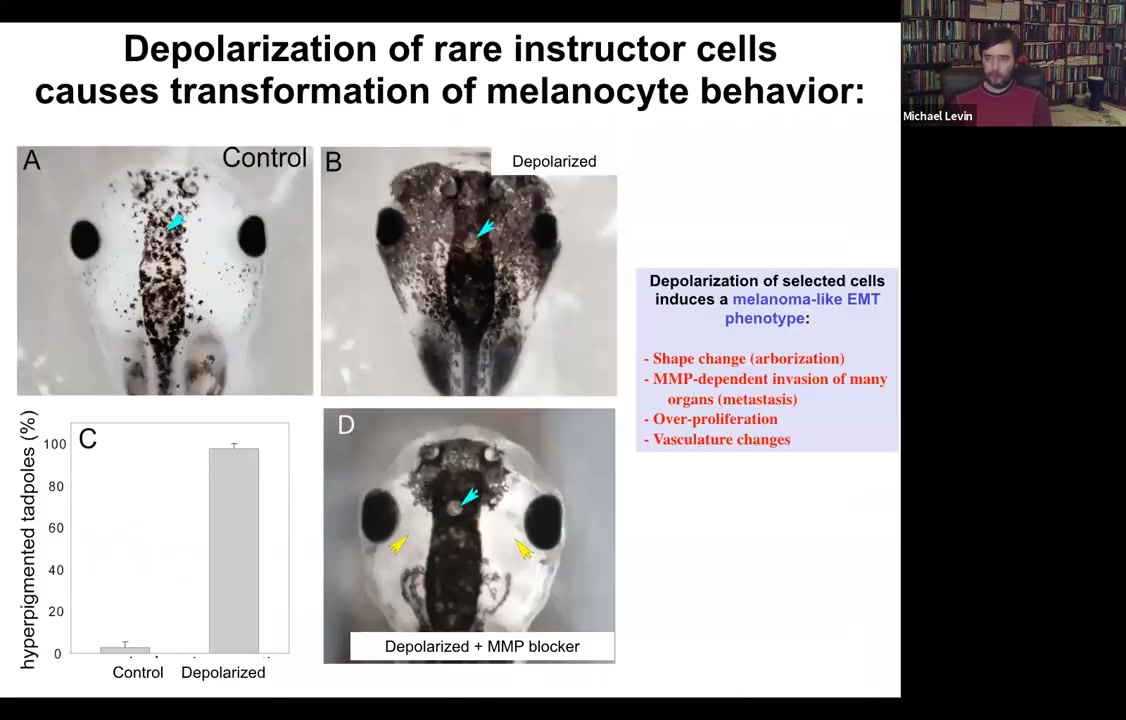
Functionally, what's really important is that you can induce a conversion of normal melanocytes. So here they are. These little black dots are pigment cells; they're melanocytes. If you depolarize a very specific cell population in the animal, not the melanocytes themselves, but a different cell population, we call them instructor cells. Those cells tell the melanocytes what to do, in particular to stay nice, cooperative melanocytes under control. If you block their ability to signal, what happens is metastatic melanoma.
Slide 46/54 · 47m:47s
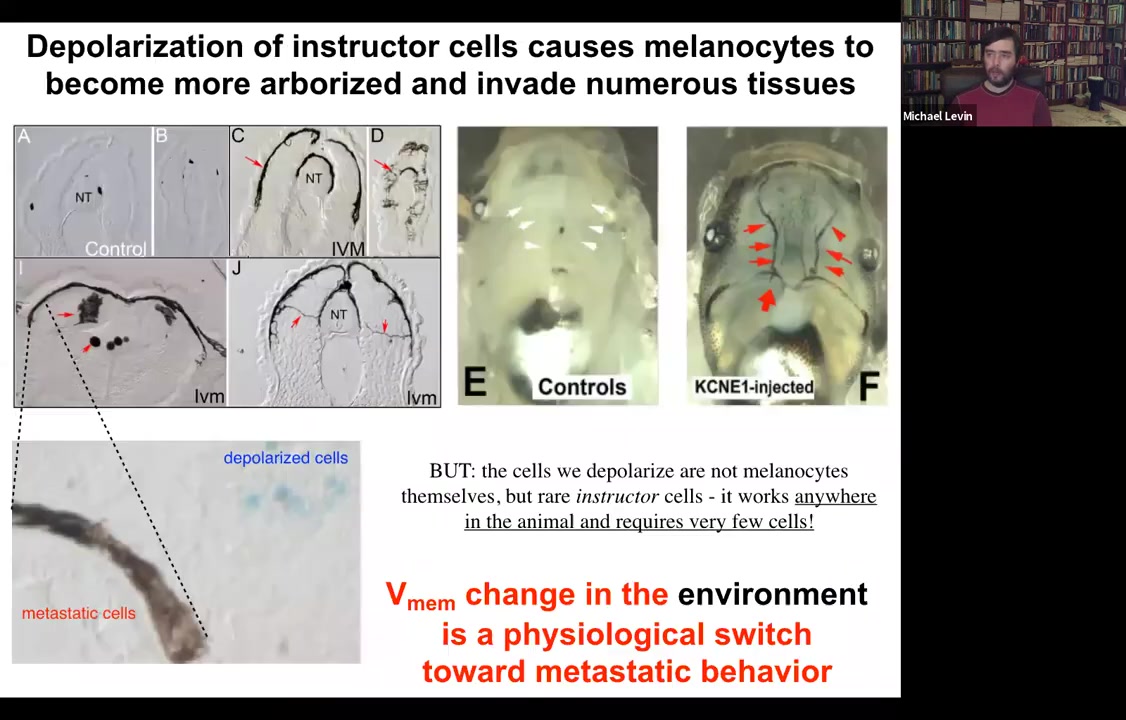
These normal melanocytes convert to these crazy, long stringy things that start to crawl away. They dig into the brain, they dig into the neural tube, they start to invade. Here they are all through the blood vessels. This is like a melanoma-type behavior. You can see here, these blue cells, these are the ones that we actually depolarized. They are the ones that are now failing to keep the normal melanocytes behaving correctly. The melanocytes are going on their own.
So it's a voltage change in the environment. There's nothing genetically wrong with these animals. There are no oncogenes, no carcinogens. But this voltage change is a physiological switch away from the cooperation of cells toward embryogenesis and toward single-cell behavior.
Slide 47/54 · 48m:44s
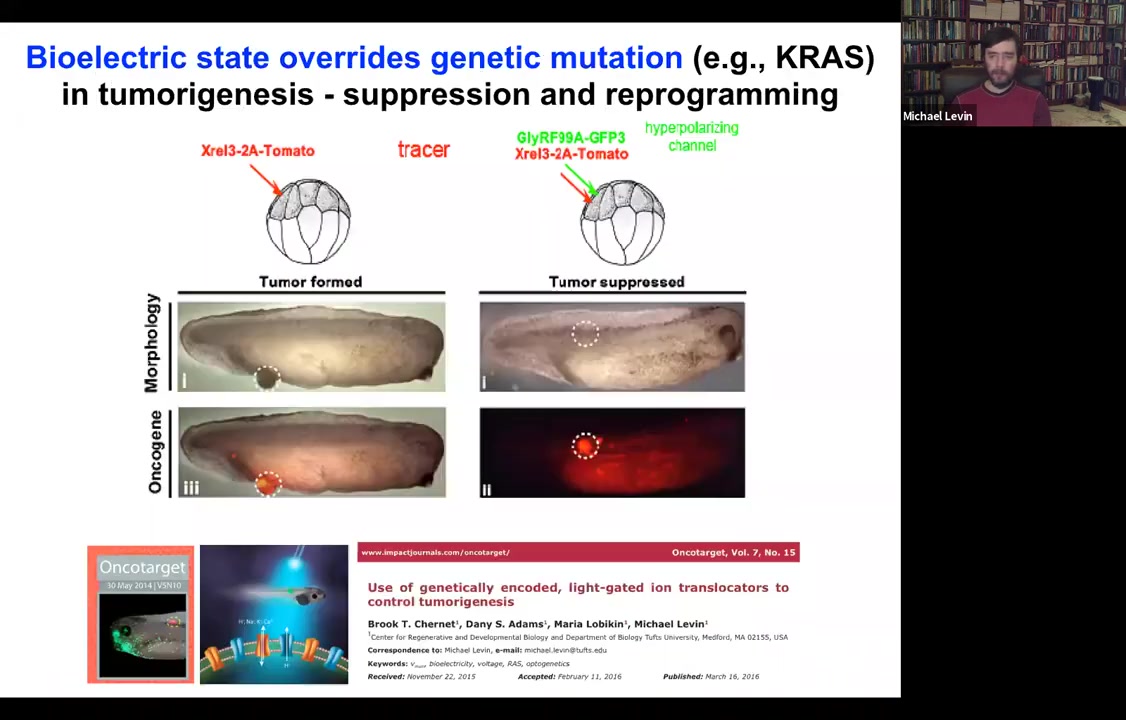
You can override this. If we inject an oncogene and they make these tumors, they're labeled with the red fluorescent protein. You can co-inject a channel and force these cells to stay electrically coupled to their neighbors despite what the oncogene is trying to get them to do.
Dominant negative KRAS, p53, while blazingly expressed, there's no tumor. It's the same animal. There's no tumor because these cells are coupled to this electrical network that forces them towards a proper tissue fate.
That's where our cancer applications are going: we've done this with light, with optogenetics, and now we're in human cells to try to use this to target carcinogenic behavior.
Slide 48/54 · 49m:34s
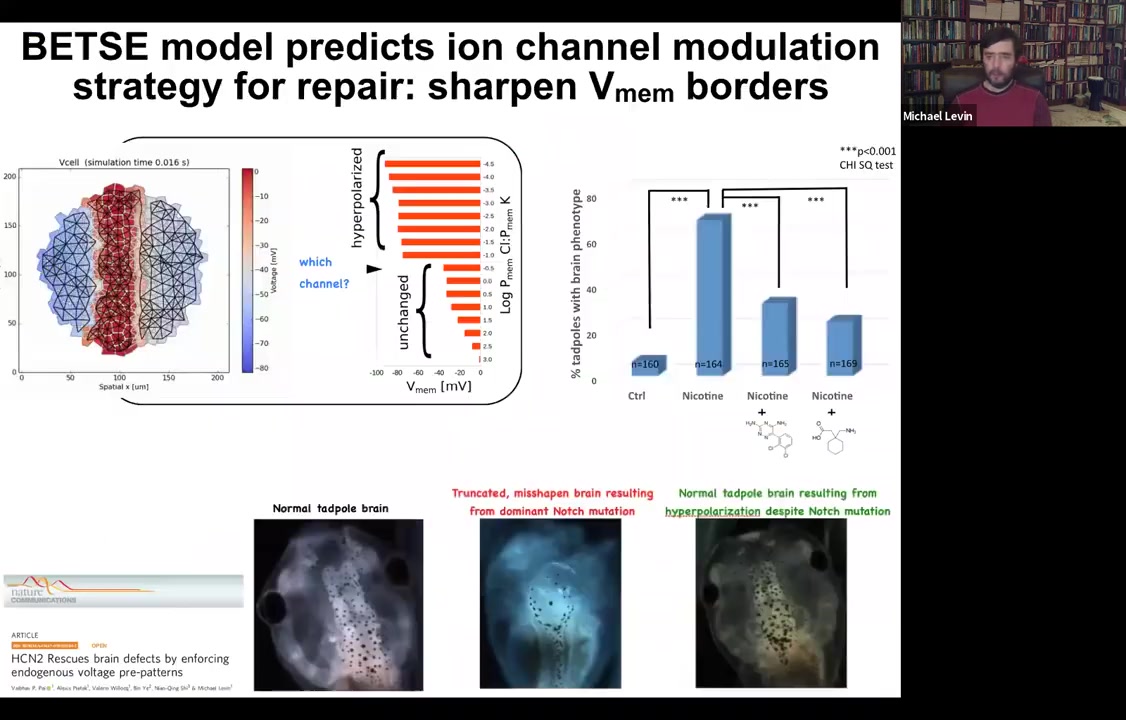
The other thing that is important is to think about the level of control that's possible with this. Here is a normal tadpole brain. You see forebrain, midbrain, and hindbrain. If you, for example, introduce a mutated Notch protein. Notch is a very important neurogenesis gene. You can mutate it and introduce a dominant mutation. You can see forebrain is basically gone, midbrain and hindbrain are just a bubble. These animals have no behavior to speak of. They lie there, they do nothing. You can do the same thing with alcohol, nicotine, various teratogen exposures.
You can build a computational model. This is the work of Alexis Pytak, who's a staff scientist in my group, and a few collaborators. You can build a computational model that asks a simple question: what determines the shape and size of the brain? There's a particular bioelectrical pattern that's required for the brain. On these backgrounds of teratogens or even Notch mutation, you can ask the model a question. You can say, if this pattern is disturbed, such as it is here, what channels would I have to open or close to get back to the correct pattern? The model told us there's a particular channel, HCN2, which can help us sharpen these boundaries. Even though there are massive defects in the Notch signaling and with these various other pathways, you can still reinforce these boundaries by opening HCN2. Sure enough, if you do it with drugs or just misexpress HCN2, you can get back to a normal brain shape, a normal brain gene expression, and in fact, normal IQ. These animals get their behaviors back and their learning rates back.
So here's what this is: using a very specific computational model to rationally manipulate the electrical signaling to get back to a very complex organ morphogenesis.
Slide 49/54 · 51m:32s
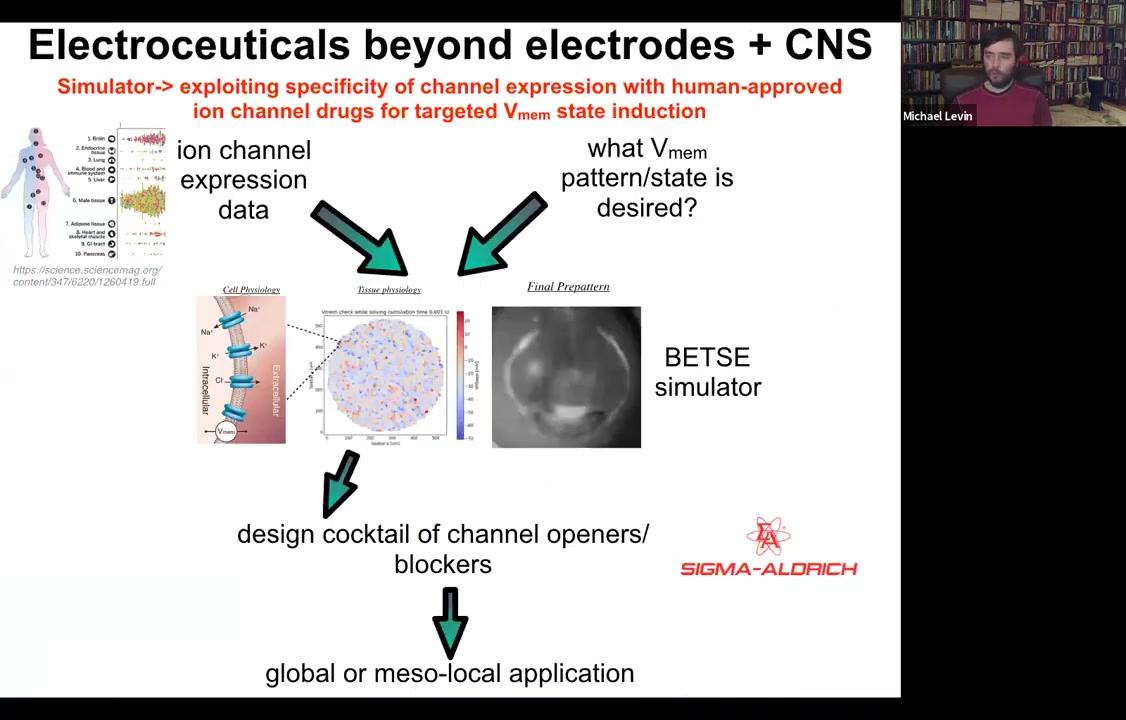
The whole roadmap looks like this. We have expression data, profiling data on which tissues in the body express which channels. That's known.
Slide 50/54 · 51m:48s
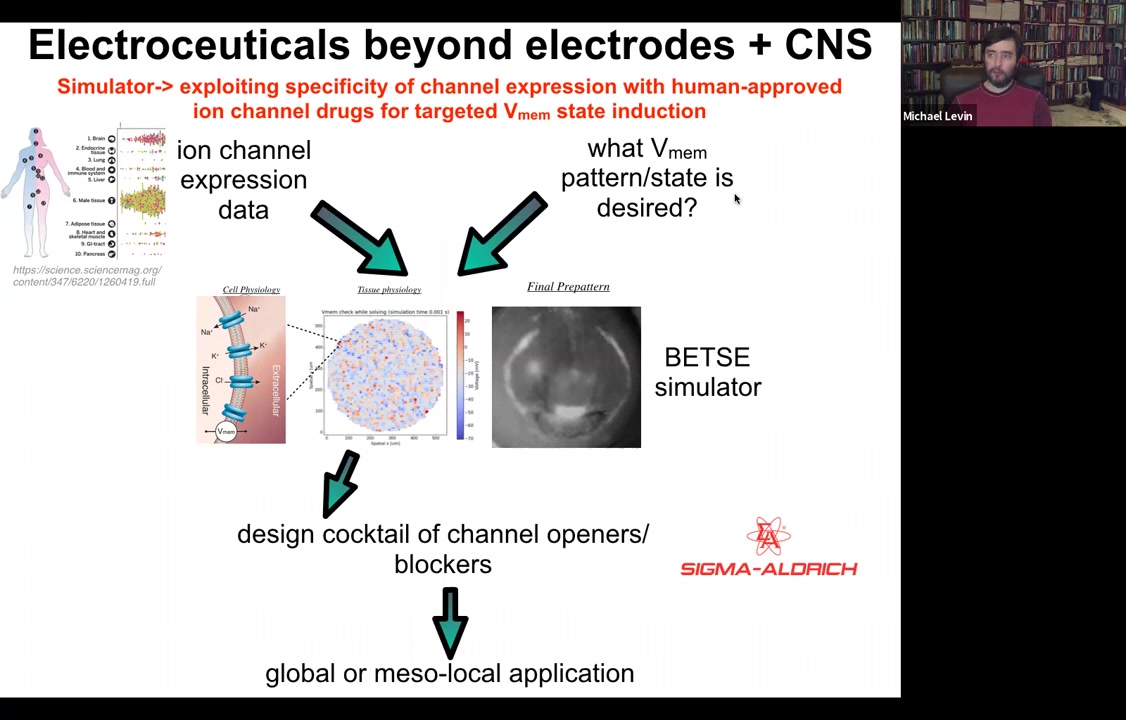
This needs a lot more work: if you knew what the voltage pattern was supposed to be in various healthy tissues under various conditions. Physiomic profiling is needed. Then we have this computational simulation engine, which is able to say, if you want to go from the incorrect pattern to the correct pattern, which of these channels you would have to open and close. That tells you immediately what cocktail of ion channel drugs you would need to use.
Slide 51/54 · 52m:16s
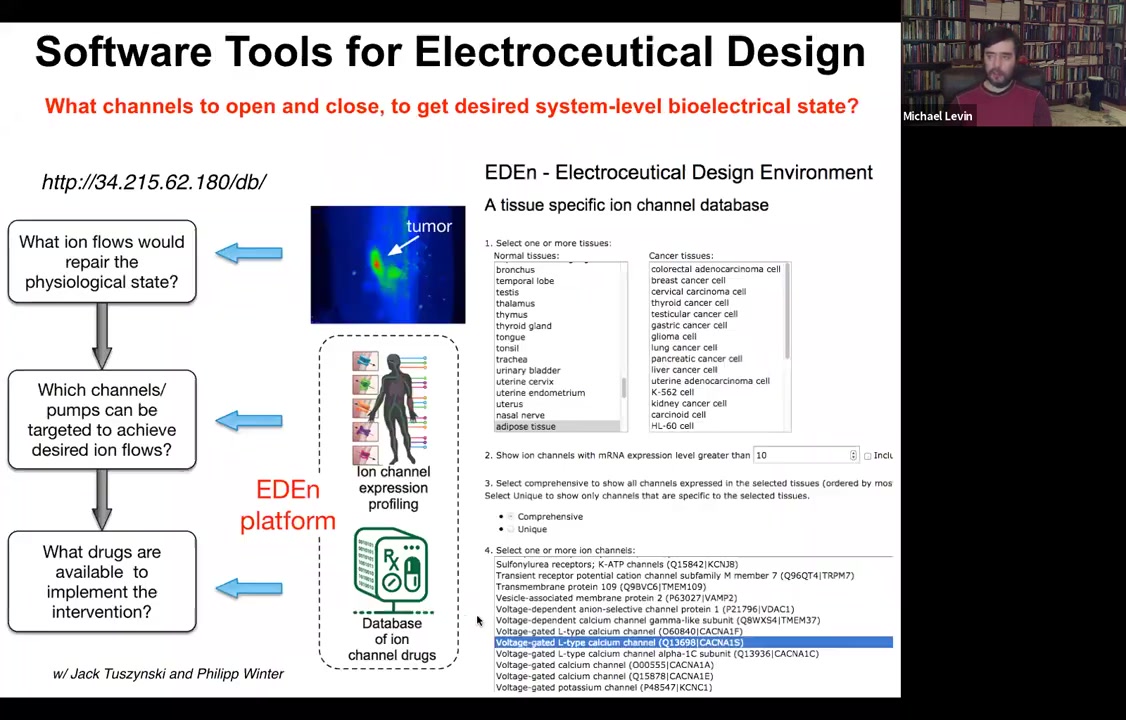

And so we've developed this pipeline. You can play with it here and choose your tissue and so on. The idea is that something like 20% of all drugs are ion channel drugs. These form an incredibly convenient toolkit of what we call electroceuticals because if you have the right computational model, you can repurpose these; the human safety data is already available. People already take them for all kinds of uses, and they have massive applicability in tweaking bioelectrical signals.
Another nice thing about this approach is that once the bioelectric circuit has made its decision, the downstream steps — second messengers, gene expression, everything else — can run as long as you want. For example, in the case of the frog, a one-day, 24-hour application of a particular cocktail gives 13 months of leg growth. You don't have to micromanage that whole process. It's just figuring out what electrical state is going to shift the tissue towards a particular goal state that all of the cells are going to work towards.
Slide 52/54 · 53m:27s
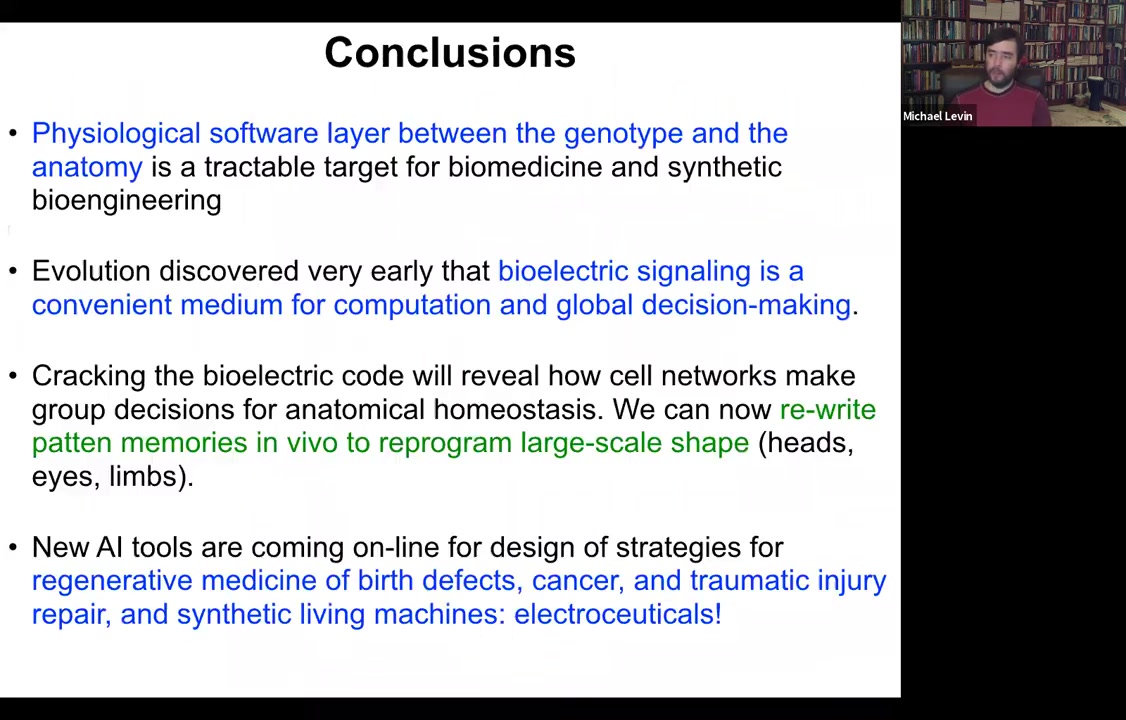

I'm going to close and summarize as follows. There's this important layer of physiological decision-making that sits between the genotype and the anatomy. It's becoming a tractable target for biomedicine and synthetic bioengineering because we can now see how the electrical dynamics operating in that tissue encode particular anatomical layouts towards which the cells continue to build.
Evolution apparently discovered very early on that this electrical signaling is a convenient medium for computation and global decision-making. It's not an accident that all nervous systems use it and all of our computer technology uses it. Ion channels, especially the voltage-gated ion channels, are basically voltage-gated current conductances. They're transistors. That's a powerful architecture that evolution found long before we did for forming feedback loops and memory circuits.
We think that cracking this bioelectric code can help reveal how cell networks make decisions in large-scale anatomy, not just how individual cells decide what type of cell they're going to be, but the real question of growth and form. Where do these complex patterns reside? We can now rewrite some of these patterns in the planarian or in the frog. New machine learning tools are coming online to help us design strategies for all kinds of applications.
Slide 53/54 · 54m:58s

I would like to thank the people in my group who did the work. These are all the postdocs and students and others who did the work that I showed you today. Here are some of our collaborators who are working with us.
The model systems that we work with are all kinds of animals. And our funders — very grateful to them. Disclosure: Morphoceuticals Inc. is the company for our frog limb regeneration work.
Slide 54/54 · 55m:26s

In the end, I always like to show a video of these two-headed animals. The first time I reported these data at a meeting, someone stood up and said, "That's impossible, those animals can't exist." Now I make sure that I bring a video.


